Richard Dawkins’s computer simulation algorithm explores how long it takes a 28-letter-long phrase to evolve to become the phrase “Methinks it is like a weasel”. The Weasel program has a single example of the phrase which produces a number of offspring, with each letter subject to mutation, where there are 27 possible letters, the 26 letters A-Z and a space. The offspring that is closest to that target replaces the single parent. The purpose of the program is to show that creationist orators who argue that evolutionary biology explains adaptations by “chance” are misleading their audiences. Pure random mutation without any selection would lead to a random sequence of 28-letter phrases. There are ![]() possible 28-letter phrases, so it should take about
possible 28-letter phrases, so it should take about ![]() different phrases before we found the target. That is without arranging that the phrase that replaces the parent is the one closest to the target. Once that highly nonrandom condition is imposed, the number of generations to success drops dramatically, from
different phrases before we found the target. That is without arranging that the phrase that replaces the parent is the one closest to the target. Once that highly nonrandom condition is imposed, the number of generations to success drops dramatically, from ![]() to mere thousands.
to mere thousands.
Although Dawkins’s Weasel algorithm is a dramatic success at making clear the difference between pure “chance” and selection, it differs from standard evolutionary models. It has only one haploid adult in each generation, and since the offspring that is most fit is always chosen, the strength of selection is in effect infinite. How does this compare to the standard Wright-Fisher model of theoretical population genetics?
The Wright-Fisher model, developed 85 years ago, independently, by two founders of theoretical population genetics, is somewhat similar. But it has an infinite number of newborns, precisely ![]() of whom survive to adulthood. In its simplest form, with asexual haploids, the adult survivors each produce an infinitely large, but equal, number of offspring. In that infinitely large offspring population, mutations occur exactly in the expected frequencies. Each genotype has a fitness, and the proportions of the genotypes change according to them, exactly as expected. Since there are an infinite number of offspring, there is no genetic drift at that stage. Then finally a random sample of
of whom survive to adulthood. In its simplest form, with asexual haploids, the adult survivors each produce an infinitely large, but equal, number of offspring. In that infinitely large offspring population, mutations occur exactly in the expected frequencies. Each genotype has a fitness, and the proportions of the genotypes change according to them, exactly as expected. Since there are an infinite number of offspring, there is no genetic drift at that stage. Then finally a random sample of ![]() of the survivors is chosen to become the adults of this new generation.
of the survivors is chosen to become the adults of this new generation.
In one case we get fairly close to the Weasel. That is when ![]() , so that there is only one adult individual. Now for some theory, part of which I explained before, in a comment here. We first assume that a match of one letter to the target multiplies the fitness by a factor of
, so that there is only one adult individual. Now for some theory, part of which I explained before, in a comment here. We first assume that a match of one letter to the target multiplies the fitness by a factor of ![]() , and that this holds no matter how many other letters match. This is the fitness scheme in the genetic algorithm program by keiths. A phrase that has
, and that this holds no matter how many other letters match. This is the fitness scheme in the genetic algorithm program by keiths. A phrase that has ![]() matches to the target has fitness
matches to the target has fitness ![]() . We also assume that mutation occurs independently in each letter of each offspring with probability
. We also assume that mutation occurs independently in each letter of each offspring with probability ![]() , and produces one of the other 26 possibilities, chosen uniformly at random. This too is what happens in keiths’s program. If a letter matches the target, there is a probability
, and produces one of the other 26 possibilities, chosen uniformly at random. This too is what happens in keiths’s program. If a letter matches the target, there is a probability ![]() that it will cease to match after mutation. If it does not match, there is a probability
that it will cease to match after mutation. If it does not match, there is a probability ![]() that it will match after mutation.
that it will match after mutation.
The interesting property of these assumptions is that, in the Wright-Fisher model with ![]() , each site evolves independently because neither its mutational process nor its selection coefficients depend on the the state of the other sites. So we can simply ask about the evolution of one site.
, each site evolves independently because neither its mutational process nor its selection coefficients depend on the the state of the other sites. So we can simply ask about the evolution of one site.
If the site matches, after mutation ![]() of the offspring will still match, and
of the offspring will still match, and ![]() of them will not. Selection results in a ratio of
of them will not. Selection results in a ratio of ![]() of matching to nonmatching letters. Dividing by the sum of these, the probability of ending up with a nonmatch is then
of matching to nonmatching letters. Dividing by the sum of these, the probability of ending up with a nonmatch is then ![]() divided by the sum, or
divided by the sum, or
![]()
Similarly if a letter does not match, after mutation the ratio of nonmatches to matches is ![]() . After selection the ratio is
. After selection the ratio is ![]() . Dividing by the sum of these, the probability of ending up with a match is
. Dividing by the sum of these, the probability of ending up with a match is
![]()
Thus, what we see at one position in the 28-letter phrase is that if it is not matched, it has probability ![]() of changing to a match, and if it is matched, it has probability
of changing to a match, and if it is matched, it has probability ![]() of changing to not being matched. This is a simple 2-state Markov process. I’m going to leave out the further analysis — I can explain any point in the comments. Suffice it to say that
of changing to not being matched. This is a simple 2-state Markov process. I’m going to leave out the further analysis — I can explain any point in the comments. Suffice it to say that
1. The probability, in the long run, of a position matching is ![]() .
.
2. This is independent in each of the 28 positions. So the number of matches will, in the long run, be a Binomial variate with 28 trials each with probability ![]() .
.
3. The “relaxation time” of this system, the number of generations to approach the long-term equilibrium, will be roughly ![]() .
.
4. At what value of ![]() do we expect selection to overcome genetic drift? There is no value where drift does not occur, but we can roughly say that when the equilibrium frequency
do we expect selection to overcome genetic drift? There is no value where drift does not occur, but we can roughly say that when the equilibrium frequency ![]() is twice as great as the no-selection value of 1/27, that selection is then having a noticeable effect. The equation for this value is a quadratic equation in
is twice as great as the no-selection value of 1/27, that selection is then having a noticeable effect. The equation for this value is a quadratic equation in ![]() whose coefficients involve
whose coefficients involve ![]() as well. If
as well. If ![]() is much smaller than
is much smaller than ![]() , then this value of
, then this value of ![]() is close to 0.44.
is close to 0.44.
5. The probability of a match at a particular position, one which started out not matching, will be after ![]() generations
generations
![]()
and the probability of a nonmatch given that one started out matching is
![]()
I don’t have the energy to see whether the runs of keiths’s program verify this, but I hope others will.

Generation: 5100 Number of survivors per generation: 1
Selection: ON Coefficient: 5.00 Mutation rate: 0.0357
Equilibrium: 70 Mean fitness: 15.61 Standard Deviation: 2.96
Making the defaults look like constants was a trick, designed to enhance usability in these particular circumstances. My sin is not to have explained the trick in a comment.
If you preserve the survivors, but convert the rest into mutated copies of the survivors, how is that non-overlapping?
// preserve the survivors, but convert the rest into mutated copies of the survivors
for (i=0; i < POPULATION_SIZE – num_survivors; i++) {
Mung,
Keep reading. The very next lines in the program answer your question:
// now mutate the survivors themselves
for (i=0; i<num_survivors; i++) {
mutate(&genome_array[i], &genome_array[i]);
}
I don’t know the genetics. But I do know a lot about computational experiments with complex dynamical systems. We have yet to investigate the impact of the finite population size on relaxation time. And even if we had observed a match to the theoretical mean value, we would want, when addressing behavior at equilibrium, to wait long enough that the probability is high that the system actually is at equilibrium. We need to wait considerably longer than the mean relaxation time to be highly confident that we’re observing the system at equilibrium. That is, the distribution of the random relaxation time has a long upper tail.
Given the uncertainties, along with the sensitivity of the mean to outliers, along with the capability to complete long runs in little time, there is no good reason to mess with data from early generations. Put simply, the data are far too inexpensive for cleverness to be justified.
Tom English,
It’s true, it was initially an idle thought in answer to an (at the time) unsaid Creationist carp about a widely variant starting population, out of which a closer-than-‘random’ approach was ‘bound’ to fall.
Tom,
Yes, and in any case Joe’s 1/(p+q) estimate is way too optimistic even with respect to the mean relaxation time.
ETA: I just did 25 runs, and in every case the relaxation time was higher than the 1/(p+q) estimate. The mean was 2.59 times the estimate.
Since it is evenhandedness, not computational efficiency, that prompted mention of equilibrium, an alternative to it would be simply to initialise the population to all X’s. Everything is then as far away as it’s possible to be – no accusations of gaming the start point – and any stochastic variation coming from the start population is also erased, by using the same start for every change of parameter.
Allan,
My soon-to-be-released new version of the program adds a command that allows you to specify a starting genotype for all members of the population.
This is a handy simplification in Markov-chain analysis. Thanks for reminding me.
I wondered whether I had misdefined relaxation time. Wikipedia on “Relaxation (physics)” says
Mine is not a continuous time process but a discrete time process with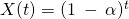 . I was saying that the relaxation time in this case would be
. I was saying that the relaxation time in this case would be 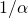 , but perhaps I should have noted that
, but perhaps I should have noted that
Then the relaxation time, defined so as to fit Wikipedia’s definition, would actually be
But that is only a little bit off from . Keiths, how are you measuring relaxation time?
. Keiths, how are you measuring relaxation time?
Joe,
I was going by your definition in the OP…
…and measuring the number of generations it took to reach the predicted equilibrium Hamming distance.
Joe Felsenstein,
I find that 100 offspring per generation, common among Weaselers (I go with considerably fewer), is not “enough” for 28 sites, 27 possible alleles at each site, mutation rate u=1/27, and selection parameter s=5. The mean observed number of fit alleles in parents over 95000 generations (in equilibrium) is generally between 15.1 and 15.4, while the Wright-Fisher model predicts 16.0.
(By the way, I’ve made an executive decision to refer to alleles that contribute to fitness as “fit alleles” instead of “matching alleles.” As you know, the math works out the same when 2 of of the 54 possible alleles at a site contribute identically to fitness as when precisely 1 of 27 contributes. But then there are 2^28 “targets” — making the term a tad silly, I hope.)
Moving up to s=100, I get a discrepancy of 0.2 (for about 1 MILLION generations). Moving up again to s=1000, the discrepancy is about 0.02. That’s practically insignificant, but “statistically significant” (a large multiple of the standard error of the sample mean ETA Oops, not large, but big enough.)
s=100.0
************************************************************************
Number of sites L : 28
Number of possible alleles A : 27
Number of offspring : 100
Mutation rate u : 0.0357142857143
Selection parameter s : 100.0
p = u * (1 + s) / (A-1 + u * s) : 0.121980676329
q = u / (1 + s * (1 – u)) : 0.000366568914956
Assume equilibrium at generation: 5000
Theoretical prob. that a site is fit p/(p+q): 0.99700386458
Expected number of fit sites Lp/(p+q) : 27.9161082083
Mean of observed numbers of fit sites : 27.7203811855
Std. dev. of observed numbers of fit sites : 0.525466911505
************************************************************************
s=1000.0
************************************************************************
Number of sites L : 28
Number of possible alleles A : 27
Number of offspring : 100
Mutation rate u : 0.0357142857143
Selection parameter s : 1000.0
p = u * (1 + s) / (A-1 + u * s) : 0.579282407407
q = u / (1 + s * (1 – u)) : 3.6998668048e-05
Assume equilibrium at generation: 5000
Theoretical prob. that a site is fit p/(p+q): 0.999936134251
Expected number of fit sites Lp/(p+q) : 27.998211759
Mean of observed numbers of fit sites : 27.9765065563
Std. dev. of observed numbers of fit sites : 0.152989617192
************************************************************************
A fine point about the monkey/Shakespeare model of cumulative selection: the “monkey” doing the imperfect copying selects on the basis of what is visible, so there’s an argument to be made that the characters are better regarded as, um, characters (traits) than as alleles. When Dawkins moves to biomorphs, he selects visible shapes, not the “genotypes” that give rise to them. His argument makes better sense if we think of the evolving sentence as a phenotype.
P.S.–I’ve modified the reporting in my code to refer to fit alleles instead of fit sites.
New code, easier to use, with detailed instructions, at Bounded Science: “Inconstant Constants to Edit, Not Mung.”
After writing all of it out, I was kind of embarrassed not to have done a graphical user interface, and prepared a proper distribution.
Mea culpa. Relaxation time in physics is defined for a deterministic process. In the present case my calculation of is actually for the change of the expectation of the match probability. That is deterministic, but the individual observations are not. Of course the time for the expectation to get all the way to the asymptote is … infinity.
is actually for the change of the expectation of the match probability. That is deterministic, but the individual observations are not. Of course the time for the expectation to get all the way to the asymptote is … infinity.
I suppose we could compute the expectation of your quantity, which is the “first passage time”. But my expected “relaxation time” is not that.
Best I could suggest in practice might be to measure the time until the number of matches gets halfway to its asymptote, then double that. That won’t exactly be the relaxation time but should be close. In a system in which departures from the equilibrium declined as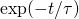 the halfway point should be approximately
the halfway point should be approximately 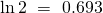 times the relaxation time. So a better estimate might be to divide the halfway time by 0.693.
times the relaxation time. So a better estimate might be to divide the halfway time by 0.693.
You’ve just been code-mined, LOL
dazz,
Leave it to Mung to invent new forms of the genre. 🙂
Joe,
I think what petrushka is really after is a way to speed up the process of getting a clean-looking histogram and accurate mean and standard deviation values. The equilibration process creates a lopsided histogram, and it takes a while for the lopsidedness to fade into the noise.
If that’s his purpose, I think his best bet is not to ignore a fixed number of generations before starting the histogram, but rather to predict the equilibrium Hamming distance and start the histogram when the program reaches that distance, which will vary from run to run In other words, always start in the middle of the histogram and let it build as the population wanders to and fro from the mean.
I didn’t have any thought out purpose.
I was responding first to the suggestion to ignore the first 2000 generations, then to the suggestion to use Joe’s formula.
My main purpose was to see if your [keiths] simulation matched Tom’s.
My untutored observation is it’s pretty close, but i wanted to match his parameters.
1000 children, 5000 generations, start computing the mean after things settle.
Right, and “after things settle” means “after equilibrium is reached”. So instead of trying to determine the settling time in advance, why not just wait until the system reaches equilibrium — whenever that happens to occur in a given run — before you start computing the mean?
It’s your program. I was a bit impatient, but I assumed you were working on the same computations. You are welcome to use anything I’ve written, if you haven’t written your own routines.
Equilibrium of the entire string of sites is equilibrium at each and every site, because the evolutionary process at each site is independent of the evolutionary processes at all other sites.
I’ve been saying that I assume equilibrium at 5000 generations. But I’m skeptical as to whether we’re observing the long-term means. I took a stab at finding finite-population results a couple days ago, but threw up my hands at it. Something that caught my eye, however, was a reference to weak convergence. I don’t know if it was in reference to a finite-population version of Wright-Fisher. But it’s highly plausible that we’re dealing with weak, and furthermore slow, convergence to the mean.
The way to start numerical experiments is with a single site. The theory essentially tells us to do that. I’ve jumped in with ad hoc results I obtained while testing my software. That’s not a proper approach to investigating the issues at hand.
petrushka,
I’ll post the current version of my code later today.
It computes the mean using the histogram data, and the histogram data starts accumulating immediately after the program starts.
If you want to speed up the process of converging toward the expected mean, you can use the ‘c’ command to clear the histogram after the program has equilibrated.
As for the ever-stinking “target” merde, I think a good response is: OK, here’s the fraction of sites that contribute (equally) to fitness in the current parent. Pick any particular string of alleles (“sentence”) you like. We’ll construct a history of particular parents consistent with the history of proportions. In fact, we can tell you the (huge) number of particular sequences of parent strings that end with one you’ve just chosen.
You see an argument that it does reach equilibrium?
Hmm….
His code has been thoroughly tested.
If I was who was referring to “weak convergence” it is relevant thusly: If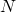 is the number of adult survivors, and
is the number of adult survivors, and  and
and  are the mutation rate and the selection coefficient, respectively, then if we take a sequence of Wright-Fisher models with
are the mutation rate and the selection coefficient, respectively, then if we take a sequence of Wright-Fisher models with 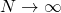 and
and 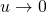 and
and  such that
such that 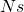 and
and 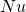 constant, and if we observe the process on a time scale where the unit is
constant, and if we observe the process on a time scale where the unit is 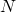 generations, the process of gene frequency change approaches a diffusion process that follows Kolmogorov’s diffusion process. The convergence of this sequence of Markov chains is technically “weak convergence”. This is the source of the rule that when
generations, the process of gene frequency change approaches a diffusion process that follows Kolmogorov’s diffusion process. The convergence of this sequence of Markov chains is technically “weak convergence”. This is the source of the rule that when 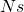 is small selection has little effect. The forces of selection, mutation, and genetic drift are all “weak” in the limit but I don’t think that this means that “weak convergence” implies that selection has little effect, or that things change slowly on the scale where one unit is a period of
is small selection has little effect. The forces of selection, mutation, and genetic drift are all “weak” in the limit but I don’t think that this means that “weak convergence” implies that selection has little effect, or that things change slowly on the scale where one unit is a period of 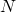 generations.
generations.
Joe Felsenstein,
Thanks. Are we on the same page? Kolmogorov backward equations (diffusion).
Joe and Keith,
Any thoughts on low-pass filters?
Tom,
Sure. The moment it arrives at the predicted equilibrium Hamming distance, it’s at equilibrium. It’s a Markov process, so the prior history is irrelevant. Only the current state matters.
I don’t have the data in front of me, but when I did my 25 runs, the average time to equilibrium was 181 generations, and the max IIRC was something like 367.
That was with s=5, u=.0357, and (I think) 5,000 offspring.
Also Kolmogorov Forward Equations, both apply to the limiting diffusion process. If you reduce each locus to a two-allele one (with alleles Match and Don’t) with appropriate mutation rates, you can find the equilibrium distribution of frequencies at that locus (position) — see Chapter VII of my online book. But the positions for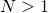 are then not really independent, but might be close. For the theory of approaches of the Wright-Fisher models to Kolmogorov diffusion processes see William Feller’s great paper in the Second Berkeley Symposium on Mathematics and Probability, available here.
are then not really independent, but might be close. For the theory of approaches of the Wright-Fisher models to Kolmogorov diffusion processes see William Feller’s great paper in the Second Berkeley Symposium on Mathematics and Probability, available here.
I’d been thinking along those lines, and went haywire above, talking half-asleep about individual sites. I apologize for adding to the sound and the fury.
The state of the process is an integer. The time-average state is a real number. So the state generally cannot match the prediction. And we’re testing the prediction, not accepting it. I’d considered averaging (many) later generations, and taking onset of equilibrium to be the first time at which the state equals the nearest integer to the average. That heuristic should work fine as long as the number of sites is not too large. It kind of bugs me, because it assumes that the time series is stationary after a point. There are statistical tests for stationarity, but it seems silly to mess with them, looking at the plots (translated: I’m lazy).
The system is amenable to analysis. I keep saying that. The fact of the matter is that I’ve spent too much time playing with my software. I’ve got an icky hacker feeling. I just downloaded QT Creator, to see if I can develop portable GUI’s quickly. So I’ll be having that icky feeling a while longer.
I’m confused, because those are for continuous time.
Be not confused. The diffusion process is the asymptotic process approached when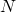 becomes infinite with
becomes infinite with 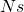 and
and 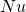 being held constant. But also, note that the time scale observed changes with
being held constant. But also, note that the time scale observed changes with 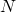 . 1 unit of the new time is
. 1 unit of the new time is 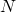 generations. So as
generations. So as 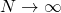 the time scale gets more and more finely divided. In the limit there is a continuous time process.
the time scale gets more and more finely divided. In the limit there is a continuous time process.
Got it. I’ve read only the introduction of Feller. It’s a good reference. Thanks for that.
Tom,
Sure, but the process will never come to rest at a non-integer Hamming distance. It’s inherently discrete. The smart choice is to use the integer closest to the predicted mean as our equilibrium criterion. Since we’re dealing with a history-independent Markov process, we’ll never be closer to equilibrium than we are the very first time we reach that Hamming distance.
Without predicting the equilibrium point, one could assume it when the SD stops falling. You might waste a few generations, but not a lot of time.
??
petrushka,
It might be quicker (I don’t know, I’m guessing) if you start with a clonal genome and assume equilibrium when the SD stops rising.
Rising schmising. Brain fart.
If you set keiths’ program to refresh the histogram on every generation, it looks quite different.
petrushka,
No, either is good. Start with a randomly variant population and let SD fall, or start with an invariant population and let it rise.
A more basic criticism of genetic algorithms is that it is very hard (read impossible) to analyze the behaviour of the GA. We expect that the mean fitness of the population will increase until an equilibrium of some kind is reached. This equilibrium is between the selection operator, which makes the population less diverse, but increases the mean fitness (exploitation), and the genetic operators, which usually reduce the mean fitness, but increase the diversity in the population (exploration). However, proving that this is guaranteed to happen has not been possible so far, which means that we cannot guarantee that the algorithm will converge at all, and certainly not to the optimal solution. This bothers a lot of researchers. That said, genetic algorithms are widely used when other methods do not work, and they are usually treated like a black box – strings are pushed in one end, and eventually an answer emerges. This is risky, because without knowledge of how the algorithm works it is not possible to improve it, nor do you know how cautiously you should treat tthe results.
– Stephen Marsland. Machine Learning: An Algorithmic Perspective
Black box huh. I should code a black box so I can learn. 🙂
Weren’t you just saying that GA’s are totally directed by the programmers input? How come the output can be so unpredictable then?
Mung,
Congratulations. You’ve mined a quote containing three substantive errors.
Last time I checked out the state of the theory of evolutionary computation (a couple years ago), the most remarkable aspect of it was that no analysis had established when there is an advantage (in optimization) of having more than one survivor per generation.
ETA: I should mention that “advantage” means a reduction in the number of fitness evaluations, not the number of generations.
All else being equal, the population genetic theory would predict greater effect of selection if the effective population size is larger. However that would be holding the strength of natural selection constant. My postdoc advisor Alan Robertson showed in 1960 that when an animal or plant breeder breeds by saving a given fraction of the distribution of offspring, where those with highest values of the trait are chosen, the optimum fraction to save is 50%, when the long-term response to selection is predicted.
That optimum comes from a tradeoff between effective population size and strength of selection. Breeding from fewer individuals lowers the former and increases the latter.