Richard Dawkins’s computer simulation algorithm explores how long it takes a 28-letter-long phrase to evolve to become the phrase “Methinks it is like a weasel”. The Weasel program has a single example of the phrase which produces a number of offspring, with each letter subject to mutation, where there are 27 possible letters, the 26 letters A-Z and a space. The offspring that is closest to that target replaces the single parent. The purpose of the program is to show that creationist orators who argue that evolutionary biology explains adaptations by “chance” are misleading their audiences. Pure random mutation without any selection would lead to a random sequence of 28-letter phrases. There are ![]() possible 28-letter phrases, so it should take about
possible 28-letter phrases, so it should take about ![]() different phrases before we found the target. That is without arranging that the phrase that replaces the parent is the one closest to the target. Once that highly nonrandom condition is imposed, the number of generations to success drops dramatically, from
different phrases before we found the target. That is without arranging that the phrase that replaces the parent is the one closest to the target. Once that highly nonrandom condition is imposed, the number of generations to success drops dramatically, from ![]() to mere thousands.
to mere thousands.
Although Dawkins’s Weasel algorithm is a dramatic success at making clear the difference between pure “chance” and selection, it differs from standard evolutionary models. It has only one haploid adult in each generation, and since the offspring that is most fit is always chosen, the strength of selection is in effect infinite. How does this compare to the standard Wright-Fisher model of theoretical population genetics?
The Wright-Fisher model, developed 85 years ago, independently, by two founders of theoretical population genetics, is somewhat similar. But it has an infinite number of newborns, precisely ![]() of whom survive to adulthood. In its simplest form, with asexual haploids, the adult survivors each produce an infinitely large, but equal, number of offspring. In that infinitely large offspring population, mutations occur exactly in the expected frequencies. Each genotype has a fitness, and the proportions of the genotypes change according to them, exactly as expected. Since there are an infinite number of offspring, there is no genetic drift at that stage. Then finally a random sample of
of whom survive to adulthood. In its simplest form, with asexual haploids, the adult survivors each produce an infinitely large, but equal, number of offspring. In that infinitely large offspring population, mutations occur exactly in the expected frequencies. Each genotype has a fitness, and the proportions of the genotypes change according to them, exactly as expected. Since there are an infinite number of offspring, there is no genetic drift at that stage. Then finally a random sample of ![]() of the survivors is chosen to become the adults of this new generation.
of the survivors is chosen to become the adults of this new generation.
In one case we get fairly close to the Weasel. That is when ![]() , so that there is only one adult individual. Now for some theory, part of which I explained before, in a comment here. We first assume that a match of one letter to the target multiplies the fitness by a factor of
, so that there is only one adult individual. Now for some theory, part of which I explained before, in a comment here. We first assume that a match of one letter to the target multiplies the fitness by a factor of ![]() , and that this holds no matter how many other letters match. This is the fitness scheme in the genetic algorithm program by keiths. A phrase that has
, and that this holds no matter how many other letters match. This is the fitness scheme in the genetic algorithm program by keiths. A phrase that has ![]() matches to the target has fitness
matches to the target has fitness ![]() . We also assume that mutation occurs independently in each letter of each offspring with probability
. We also assume that mutation occurs independently in each letter of each offspring with probability ![]() , and produces one of the other 26 possibilities, chosen uniformly at random. This too is what happens in keiths’s program. If a letter matches the target, there is a probability
, and produces one of the other 26 possibilities, chosen uniformly at random. This too is what happens in keiths’s program. If a letter matches the target, there is a probability ![]() that it will cease to match after mutation. If it does not match, there is a probability
that it will cease to match after mutation. If it does not match, there is a probability ![]() that it will match after mutation.
that it will match after mutation.
The interesting property of these assumptions is that, in the Wright-Fisher model with ![]() , each site evolves independently because neither its mutational process nor its selection coefficients depend on the the state of the other sites. So we can simply ask about the evolution of one site.
, each site evolves independently because neither its mutational process nor its selection coefficients depend on the the state of the other sites. So we can simply ask about the evolution of one site.
If the site matches, after mutation ![]() of the offspring will still match, and
of the offspring will still match, and ![]() of them will not. Selection results in a ratio of
of them will not. Selection results in a ratio of ![]() of matching to nonmatching letters. Dividing by the sum of these, the probability of ending up with a nonmatch is then
of matching to nonmatching letters. Dividing by the sum of these, the probability of ending up with a nonmatch is then ![]() divided by the sum, or
divided by the sum, or
![]()
Similarly if a letter does not match, after mutation the ratio of nonmatches to matches is ![]() . After selection the ratio is
. After selection the ratio is ![]() . Dividing by the sum of these, the probability of ending up with a match is
. Dividing by the sum of these, the probability of ending up with a match is
![]()
Thus, what we see at one position in the 28-letter phrase is that if it is not matched, it has probability ![]() of changing to a match, and if it is matched, it has probability
of changing to a match, and if it is matched, it has probability ![]() of changing to not being matched. This is a simple 2-state Markov process. I’m going to leave out the further analysis — I can explain any point in the comments. Suffice it to say that
of changing to not being matched. This is a simple 2-state Markov process. I’m going to leave out the further analysis — I can explain any point in the comments. Suffice it to say that
1. The probability, in the long run, of a position matching is ![]() .
.
2. This is independent in each of the 28 positions. So the number of matches will, in the long run, be a Binomial variate with 28 trials each with probability ![]() .
.
3. The “relaxation time” of this system, the number of generations to approach the long-term equilibrium, will be roughly ![]() .
.
4. At what value of ![]() do we expect selection to overcome genetic drift? There is no value where drift does not occur, but we can roughly say that when the equilibrium frequency
do we expect selection to overcome genetic drift? There is no value where drift does not occur, but we can roughly say that when the equilibrium frequency ![]() is twice as great as the no-selection value of 1/27, that selection is then having a noticeable effect. The equation for this value is a quadratic equation in
is twice as great as the no-selection value of 1/27, that selection is then having a noticeable effect. The equation for this value is a quadratic equation in ![]() whose coefficients involve
whose coefficients involve ![]() as well. If
as well. If ![]() is much smaller than
is much smaller than ![]() , then this value of
, then this value of ![]() is close to 0.44.
is close to 0.44.
5. The probability of a match at a particular position, one which started out not matching, will be after ![]() generations
generations
![]()
and the probability of a nonmatch given that one started out matching is
![]()
I don’t have the energy to see whether the runs of keiths’s program verify this, but I hope others will.

This is average number of matches, right?
Keiths uses 50000.
I can change it.
That’s the way I read it.
I took one term of statistics in 1970, and haven’t seen a C compiler since 1995.
I’m a bit under-educated and rusty.
With 1000 children per generation:
Generation: 7040 Number of survivors per generation: 1
Selection: ON Coefficient: 5.00 Mutation rate: 0.036
Mean fitness: 16.04 Standard Deviation: 2.78
Another run:
Mean fitness: 15.59 Standard Deviation: 2.65
and another:
Mean fitness: 15.68 Standard Deviation: 2.87
Doing this with the data, again, from the red run above, I get a mean of 15.934730538922155 and a standard deviation of 2.575163769620239. The histogram isn’t so pretty this time, so I probably was just lucky with the black one above.
Keiths will have to look at my SD code in the context of his program and tell us if it is done correctly, and if I’m using the right snapshot of his run.
This is looking reasonable. The standard deviation is telling you to do some longer runs, if you want to see if the mean observation is close to the theoretical mean.
Joe’s interested also in the time for “relaxation,” and that will take many short runs (but I would rather take a shot at analysis before getting into that).
Joe pointed out that I shouldn’t have called it fitness, and he was right.
Joe pointed out that I shouldn’t have called it fitness, and he was right.
Oh, I thought petrushka was running keiths’ algo, not yours, and I also got lost in translation and somehow thought mean = median. I’m on a roll, duh!
Thanks. Implementing it in a different language is a good way to study it.
I am running keiths’ program. I just added code to calculate the mean and sd. Saves time comparing it with Tom’s.
I just added the caveat that I am not a wizard. I’m pretty sure I got it right, but I wouldn’t ship it.
Do you have a windows build please? I can’t get it to compile on MinGW
http://www.filedropper.com/keithssd
This is one I modified to display mean and sd.
It is also modified to default to 1000 children and a mutation rate of .0357.
These can be changed interactively.
petrushka,
Thank you! I almost have the code compiled in MinGW, apparently conio is not available in it’s compiler :/ Might just remove that even if I can’t change params on the fly
It’s possible that conio.lib might sneak it’s way to your computer. If you wished it.
Lcc-win is free and takes about five minutes to download and install.
Keiths got it working on Win-7.
I have it working on win-10 and XP.
Thanks
On Windows you might try installing Git for Windows and running from a git bash prompt.
ETA: Nope.
ks-drift-weasel.c:6:24: fatal error: sys/select.h: No such file or directory
ks-drift-weasel.c:7:21: fatal error: termios.h: No such file or directory
I’ll be out for the evening, leaving in a while. I think I’ll leave a download for my sd modification. I did not rename it, so it might sit on top of any previous code.
http://www.filedropper.com/keiths
This is keiths.c
Those are the linux i/o libs. Might still compile with petrushka’s tweaks
http://pastebin.com/raw/hE14eHGN
Thanks for the suggestion, I’m rather incompetent in C
I didn’t miss the sarcasm.
Are there any such critics of evolutionary theory who still think that it is composed only of chance mechanisms or would descibe it as such?
Yes of course its not random. I’m making an observation that Weasel is modelling a process that has a target to aim at.
And as I see that Mung has started a thread to prevent the diversions that my post will inevitably bring if I keep getting responses, I’m happy to continue over there.
I’m back,if anyone cares. The more recent versions of keiths’ code will compile without file errors, but you need to edit the dot c file and add a line at the top that says #define WINDOWS
Here I think is where the ambiguity sets in. In the Dawkins example, the exact target is specified at the start, and all iterations are compared with that target. This doesn’t resemble evolution, it’s simply an illustration of the power of selection. The question is, if the “target” were more general, like “produce a grammatically correct sentence” would that count as a pre-specified target?
After all, biological evolution also has a pre-specified target in this sense – produce an organism capable of surviving long enough to breed in any environment available to be colonized.
In the Joe F model, the target is irrelevant except as a way of measuring drift. It’s just a mathematical abstraction. What’s important is the mathematics of the selection coefficient.
lcc worked like a charm, thanks again
petrushka,
Strictly speaking, the number discarded should be variable, based on s and u – according to Joe’s OP point (3), equilibrium takes 1/(p+q) generations, with p and q both having u and s terms.
That is food for thought. This would give the Weasel algorithm an unimaginable number of targets to aim for, not just one pre-specified target.
Can the Weasel algorithm be adjusted to test this? Has it ever been tried previously?
The GA to calculate PI by combining digits and arithmetic symbols did exactly that. There are literally an infinitude of possible arithmetic expressions, and since PI is infinite it’s impossible to reach the “target”, just find better and better solutions
Dawkins, after presenting his “Weasel” program on pp 46 – 50 of Blind Watchmaker, went on to describe ‘Biomorph Land’, his attempt to develop a more realistic computer model of selection. In one way, it is a pretty good model of artificial (or indeed, sexual) selection, with the operator playing the rôle of God, the environment, selective breeder or choosy mate.
I have a web game in which the “target” is all the words in the Scrabble dictionary. One of the regulars at AtBC did something similar.
Not an unimaginable number, but over a hundred thousand possible targets. Works in five languages. Glad you asked.
The more targets, the better.
I’ve done something wrong. Using mutation rate of .0357 and selection coefficient of 5, I get a relaxation time of 69.
cs = CHARSET_SIZE - 1;
u = mutation_rate;
s = selection_coefficient;
q = u / (1 + (s * (1 - u)));
p = (u * (1 + s)) / (cs + (u * s));
eq = 1 / (p + q);
Holy shit, I see the stupidity here hasn’t abaded one bit.
A program, which has in place a solution (a never changing solution!), and which simply picks things that match that solution, is an example of how evolution is not random.
Fucking hell. What a waste of an education.
phoodoo,
Let us not detain you further, then.
petrushka,
It may be the units for ‘generations’. 5000 births/deaths in a steady state population of 1000 would be 5 generations (phoodoo’s back to set that one straight!).
In keiths’ code, a generation is independent of child population size.
First statement, first blunder. Pathetic
Hi Tom,
Was it your intent to have this refer to your constants at the top of the file?
Evolve(n_offspring=1000, selection=s, alphabet_size=27)
Line 100
ETA: ah. I think I see you are just using the constants for defaults.
I am so glad that Mung started a new thread so that all the meaningless baiting about “it’s a search” and “it has a target built in” can happen there and not here …
For those who want to see an evolutionary simulation which does not have a target built in, there are genetic algorithms that do not have any detailed specification of the goal genotype or phenotype built in, but achieve remarkable adaptations:
(1) Karl Sims (site here) had a genetic algorithm that described organisms made of blocks and links that existed in a simulated physics. They were under selection (to pick a dramatic case) to swim through the medium to the right. Starting with phenotypes that could barely wiggle, they evolve to “swim” in all sorts of fascinating ways. Sims’s code is not available but there is an imitation of it called
brevethat is available from Jonathan Klein (here).(2) A similar effort called
Boxcar2devolves imaginary vehicles that move to the right across a 2-dimensional landscape. It will be found here.I have mentioned these here before.
Joe Felsenstein,
How well did the boxcar2d algorithm do at creating a better antibiotic for staph infection?
I am guessing not so good. Funny how all these so called random algorithms only do the thing they are supposed to do.
Gee, I wonder why people think they are searches Joe?
phoodoo,
Indeed, you seem to carry it with you wherever you go.
Perhaps cracking a book or two would help ‘abade’ it.
Your ETA is correct. It was a lapse in judgment. I was shooting for a combination of ease of use and compact code, when I should have favored flexibility and explicitness in use. I’ll post a no-constants version on my blog, and send you a PM.
petrushka:
Allan:
Strictly speaking, you don’t have to discard anything. The mean will approach the ‘true’ value over time anyway. Discarding the initial generations just speeds up the process somewhat.
I implemented the ‘c’ command — the one that resets the histogram — as a convenience, not as a necessity.
Allan:
petrushka:
That’s right. The generations are non-overlapping, which is why the Wright-Fisher model is applicable.
Hi Tom.
I like being able to change the settings at the top of the file but if the program uses the constants then there’s not much point in setting defaults, lol. Whatever you come up with I’ll try not to bitch about it because off the cuff I couldn’t think of a better way. 🙂
Thanks. Now if folks can just resist the temptation to respond to baiting in this thread. Please.
Joe:
And as dazz mentioned, the GA in this thread also qualifies:
Math Genome Fun 2, UPB and beyond?
The solutions the GA comes up with are completely different depending on the constant being approximated and the digits available to the GA. It’s obvious that the solutions are not being “smuggled” in.
maybe you could take a look at this and see if anything’s wrong.
http://www.filedropper.com/keiths-petru
http://www.labbookpages.co.uk/teaching/evoHW/files/ehw3.pdf
Look at Thompson’s tone discriminator.
petrushka,
Are you asking about the relaxation time computation?
If so, I think your code is correct, but I don’t know the basis of the 1/(p+q) formula. Joe, could you elaborate?
I showed in the OP my formulas for the probability of change from nonmatch to match in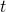 generations. Basically the probability of match rises from
generations. Basically the probability of match rises from 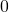 to
to 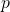 in the first generation. The asymptotic value is
in the first generation. The asymptotic value is 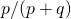 . Thus in one generation it goes a fraction
. Thus in one generation it goes a fraction
of the way to that asymptote. If it continued changing by the amount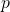 it would then reach the asymptote in
it would then reach the asymptote in  generations. Of course, it doesn’t do that, it changes by less as it approaches the asymptote. But the
generations. Of course, it doesn’t do that, it changes by less as it approaches the asymptote. But the  time is a reasonable indication of how fast it is changing.
time is a reasonable indication of how fast it is changing.
A similar calculation can be done for the case where we start with a match, It gives the same answer for the relaxation time.