Richard Dawkins’s computer simulation algorithm explores how long it takes a 28-letter-long phrase to evolve to become the phrase “Methinks it is like a weasel”. The Weasel program has a single example of the phrase which produces a number of offspring, with each letter subject to mutation, where there are 27 possible letters, the 26 letters A-Z and a space. The offspring that is closest to that target replaces the single parent. The purpose of the program is to show that creationist orators who argue that evolutionary biology explains adaptations by “chance” are misleading their audiences. Pure random mutation without any selection would lead to a random sequence of 28-letter phrases. There are ![]() possible 28-letter phrases, so it should take about
possible 28-letter phrases, so it should take about ![]() different phrases before we found the target. That is without arranging that the phrase that replaces the parent is the one closest to the target. Once that highly nonrandom condition is imposed, the number of generations to success drops dramatically, from
different phrases before we found the target. That is without arranging that the phrase that replaces the parent is the one closest to the target. Once that highly nonrandom condition is imposed, the number of generations to success drops dramatically, from ![]() to mere thousands.
to mere thousands.
Although Dawkins’s Weasel algorithm is a dramatic success at making clear the difference between pure “chance” and selection, it differs from standard evolutionary models. It has only one haploid adult in each generation, and since the offspring that is most fit is always chosen, the strength of selection is in effect infinite. How does this compare to the standard Wright-Fisher model of theoretical population genetics?
The Wright-Fisher model, developed 85 years ago, independently, by two founders of theoretical population genetics, is somewhat similar. But it has an infinite number of newborns, precisely ![]() of whom survive to adulthood. In its simplest form, with asexual haploids, the adult survivors each produce an infinitely large, but equal, number of offspring. In that infinitely large offspring population, mutations occur exactly in the expected frequencies. Each genotype has a fitness, and the proportions of the genotypes change according to them, exactly as expected. Since there are an infinite number of offspring, there is no genetic drift at that stage. Then finally a random sample of
of whom survive to adulthood. In its simplest form, with asexual haploids, the adult survivors each produce an infinitely large, but equal, number of offspring. In that infinitely large offspring population, mutations occur exactly in the expected frequencies. Each genotype has a fitness, and the proportions of the genotypes change according to them, exactly as expected. Since there are an infinite number of offspring, there is no genetic drift at that stage. Then finally a random sample of ![]() of the survivors is chosen to become the adults of this new generation.
of the survivors is chosen to become the adults of this new generation.
In one case we get fairly close to the Weasel. That is when ![]() , so that there is only one adult individual. Now for some theory, part of which I explained before, in a comment here. We first assume that a match of one letter to the target multiplies the fitness by a factor of
, so that there is only one adult individual. Now for some theory, part of which I explained before, in a comment here. We first assume that a match of one letter to the target multiplies the fitness by a factor of ![]() , and that this holds no matter how many other letters match. This is the fitness scheme in the genetic algorithm program by keiths. A phrase that has
, and that this holds no matter how many other letters match. This is the fitness scheme in the genetic algorithm program by keiths. A phrase that has ![]() matches to the target has fitness
matches to the target has fitness ![]() . We also assume that mutation occurs independently in each letter of each offspring with probability
. We also assume that mutation occurs independently in each letter of each offspring with probability ![]() , and produces one of the other 26 possibilities, chosen uniformly at random. This too is what happens in keiths’s program. If a letter matches the target, there is a probability
, and produces one of the other 26 possibilities, chosen uniformly at random. This too is what happens in keiths’s program. If a letter matches the target, there is a probability ![]() that it will cease to match after mutation. If it does not match, there is a probability
that it will cease to match after mutation. If it does not match, there is a probability ![]() that it will match after mutation.
that it will match after mutation.
The interesting property of these assumptions is that, in the Wright-Fisher model with ![]() , each site evolves independently because neither its mutational process nor its selection coefficients depend on the the state of the other sites. So we can simply ask about the evolution of one site.
, each site evolves independently because neither its mutational process nor its selection coefficients depend on the the state of the other sites. So we can simply ask about the evolution of one site.
If the site matches, after mutation ![]() of the offspring will still match, and
of the offspring will still match, and ![]() of them will not. Selection results in a ratio of
of them will not. Selection results in a ratio of ![]() of matching to nonmatching letters. Dividing by the sum of these, the probability of ending up with a nonmatch is then
of matching to nonmatching letters. Dividing by the sum of these, the probability of ending up with a nonmatch is then ![]() divided by the sum, or
divided by the sum, or
![]()
Similarly if a letter does not match, after mutation the ratio of nonmatches to matches is ![]() . After selection the ratio is
. After selection the ratio is ![]() . Dividing by the sum of these, the probability of ending up with a match is
. Dividing by the sum of these, the probability of ending up with a match is
![]()
Thus, what we see at one position in the 28-letter phrase is that if it is not matched, it has probability ![]() of changing to a match, and if it is matched, it has probability
of changing to a match, and if it is matched, it has probability ![]() of changing to not being matched. This is a simple 2-state Markov process. I’m going to leave out the further analysis — I can explain any point in the comments. Suffice it to say that
of changing to not being matched. This is a simple 2-state Markov process. I’m going to leave out the further analysis — I can explain any point in the comments. Suffice it to say that
1. The probability, in the long run, of a position matching is ![]() .
.
2. This is independent in each of the 28 positions. So the number of matches will, in the long run, be a Binomial variate with 28 trials each with probability ![]() .
.
3. The “relaxation time” of this system, the number of generations to approach the long-term equilibrium, will be roughly ![]() .
.
4. At what value of ![]() do we expect selection to overcome genetic drift? There is no value where drift does not occur, but we can roughly say that when the equilibrium frequency
do we expect selection to overcome genetic drift? There is no value where drift does not occur, but we can roughly say that when the equilibrium frequency ![]() is twice as great as the no-selection value of 1/27, that selection is then having a noticeable effect. The equation for this value is a quadratic equation in
is twice as great as the no-selection value of 1/27, that selection is then having a noticeable effect. The equation for this value is a quadratic equation in ![]() whose coefficients involve
whose coefficients involve ![]() as well. If
as well. If ![]() is much smaller than
is much smaller than ![]() , then this value of
, then this value of ![]() is close to 0.44.
is close to 0.44.
5. The probability of a match at a particular position, one which started out not matching, will be after ![]() generations
generations
![]()
and the probability of a nonmatch given that one started out matching is
![]()
I don’t have the energy to see whether the runs of keiths’s program verify this, but I hope others will.

Mung,
No, what you should do is sneer at the very idea you could learn anything from evolutionary computing.
I don’t think it’s very smart to assume the truth of what is in question. In the figure below, the dashed line is the model prediction.
Some results using keiths’ program
Tom,
Who’s assuming?
We’re dealing with a Markov process. Joe analyzed it as one, and I implemented it as one in my program. The future behavior of a Markov process depends only on its current state, not on its prior history.
In run after run of my program, the observed mean gravitates to the predicted mean and stays there, even in runs that last for millions of generations. The histograms display the expected symmetry. The model and the program agree.
Given all of that, I stand by my statement:
Is that a new record?
Trying again
Mean fitness at generation 5000
Mutation Rate 0.0357
1000 survivors
Selection 20, survivors 1 mutation rate 0.0357
If I use a higher number of survivors, keiths’ program quickly goes to maximum fitness.
Seems to be a bug in my calculation of fitness. Possibly calculating generation zero.
We are dealing with different Markov processes, plural, than the one that Joe analyzed. The question is how well the infinite-offspring analysis applies to finite-offspring cases.
Then you’ve done little exploration of the parameter space. I provided you with parameter settings that you have not considered. It’s strange that you should ignore them, and reassert your correctness.
petrushka,
You’re evidently plotting average values, so we need to know how you’re averaging. Also, it’s as important to know the number of offspring as the number of survivors. The curve does stick close to the mean for infinitely many offspring,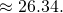 According to my direct implementation of the finite-population model, it does not for
According to my direct implementation of the finite-population model, it does not for
The wallclock time for the run generating the following four plots was 1 minute, 8 seconds. I’m holding everything but the number of offspring constant.
ETA: Tried to include all four plots at once. Didn’t work.
Number of offspring: 10
************************************************************************
Number of sites L : 28
Number of possible alleles A : 27
Number of offspring : 10
Mutation rate u : 0.037037037037
Selection parameter s : 20.0
p = u * (1 + s) / (A-1 + u * s) : 0.0290858725762
q = u / (1 + s * (1 – u)) : 0.0018281535649
Assume equilibrium at generation: 0
W-F probability of fit allele at site p/(p+q) : 0.940863297567
Expected num. of fit alleles in parent Lp/(p+q): 26.3441723319
Mean observed num. of fit alleles in parent : 19.9324506755
Std. dev. of observed numbers of fit alleles : 2.17372026547
************************************************************************
Number of offspring: 20
************************************************************************
Number of sites L : 28
Number of possible alleles A : 27
Number of offspring : 20
Mutation rate u : 0.037037037037
Selection parameter s : 20.0
p = u * (1 + s) / (A-1 + u * s) : 0.0290858725762
q = u / (1 + s * (1 – u)) : 0.0018281535649
Assume equilibrium at generation: 0
W-F probability of fit allele at site p/(p+q) : 0.940863297567
Expected num. of fit alleles in parent Lp/(p+q): 26.3441723319
Mean observed num. of fit alleles in parent : 22.800301997
Std. dev. of observed numbers of fit alleles : 1.98878876606
************************************************************************
Number of offspring: 30
************************************************************************
Number of sites L : 28
Number of possible alleles A : 27
Number of offspring : 30
Mutation rate u : 0.037037037037
Selection parameter s : 20.0
p = u * (1 + s) / (A-1 + u * s) : 0.0290858725762
q = u / (1 + s * (1 – u)) : 0.0018281535649
Assume equilibrium at generation: 0
W-F probability of fit allele at site p/(p+q) : 0.940863297567
Expected num. of fit alleles in parent Lp/(p+q): 26.3441723319
Mean observed num. of fit alleles in parent : 23.8999310007
Std. dev. of observed numbers of fit alleles : 1.85098401771
************************************************************************
Number of offspring: 40
************************************************************************
Number of sites L : 28
Number of possible alleles A : 27
Number of offspring : 40
Mutation rate u : 0.037037037037
Selection parameter s : 20.0
p = u * (1 + s) / (A-1 + u * s) : 0.0290858725762
q = u / (1 + s * (1 – u)) : 0.0018281535649
Assume equilibrium at generation: 0
W-F probability of fit allele at site p/(p+q) : 0.940863297567
Expected num. of fit alleles in parent Lp/(p+q): 26.3441723319
Mean observed num. of fit alleles in parent : 24.4291057089
Std. dev. of observed numbers of fit alleles : 1.74624338383
************************************************************************
10 offspring
20 offspring
30 offspring. The first plot in this series was 40 offspring.
Reminder: To minimize the expected number of function evaluations to hit the target in the straight Weasel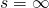 , set the number of offspring to 20, and the mutation rate to a value close to what we’re looking at here (ETA: 1/27 in my last four plots — I normally go with 1/L). See the bottom curve on the second of DiEb’s two plots here (but keep in mind that he’s giving rates of “lazy” mutation, which changes the character with probability 26/27).
, set the number of offspring to 20, and the mutation rate to a value close to what we’re looking at here (ETA: 1/27 in my last four plots — I normally go with 1/L). See the bottom curve on the second of DiEb’s two plots here (but keep in mind that he’s giving rates of “lazy” mutation, which changes the character with probability 26/27).
The earliest of my runs were with small numbers of offspring.
Tom,
Yes, and I have already investigated that via my program. My equilibrium criterion is based on both theory and observation. It’s not an assumption, contrary to your characterization.
keiths:
Tom:
Are you talking about the small-population cases? I’ve avoided those quite deliberately, because we already know they will depart from Joe’s model. I’m trying to approximate Joe’s model, not diverge from it.
Why?
Because agreement between the predictions of Joe’s model and the performance of my program increases my confidence in the correctness of both.
Joe’s model assumes an infinitude of offspring. Using a large number of offspring in my program will approximate that better than using a small number, so predictions (such as the mean Hamming distance) are more likely to match well when I use large numbers.
One should add one prediction. For the Wright-Fisher model in the case of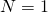 the individual positions are independent, so the distribution of matches should be a binomial distribution with 28 trials and probability
the individual positions are independent, so the distribution of matches should be a binomial distribution with 28 trials and probability 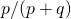 . This I mentioned before. It is rather annoying to test, but at least the variance (and the standard deviation) can be checked. If
. This I mentioned before. It is rather annoying to test, but at least the variance (and the standard deviation) can be checked. If 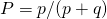 the standard deviation should be at equilibrium
the standard deviation should be at equilibrium
If that fits, the distribution wll at least have the right width.
Joe,
I’ll update the latest version of my program to display both the predicted and observed standard deviations.
Joe Felsenstein,
Let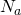 denote the size of the alphabet. It appears that you took the approximate value of
denote the size of the alphabet. It appears that you took the approximate value of 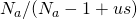 to be 1. With
to be 1. With 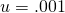 and
and 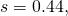 this works fine for
this works fine for 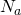 of at least 20 or so. It works poorly for
of at least 20 or so. It works poorly for 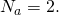
Tom English,
I didn’t test it for any value except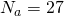 . You’d expect it to work poorly for
. You’d expect it to work poorly for 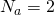 because then twice the value of
because then twice the value of 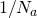 would be 1, which is as high as the equilibrium value could get. Any higher and you’re in real trouble.
would be 1, which is as high as the equilibrium value could get. Any higher and you’re in real trouble.
That’s precisely what I expected. Most of what you’ve said holds for all alphabet sizes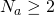 , even though you refer to 27 throughout. It appears that you’ve provided an asymptotic result that works for
, even though you refer to 27 throughout. It appears that you’ve provided an asymptotic result that works for 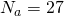 because the curve for
because the curve for 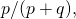 a function of
a function of 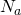 and
and  , converges on
, converges on 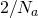 rapidly when
rapidly when  is much smaller than constant
is much smaller than constant 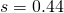 .
.
A Weasel run showing predicted vs observed mean Hamming distance and standard deviation after 800,000 generations:
Keiths, have you tested with more than one survivor?
My result is not asymptotic for large alphabets. It is exact for alphabet size but is asymptotic in the number of offspring. Of course you can replace 25, 26, and 27 by respectively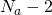 ,
, 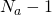 . and
. and 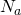 and you get the theory for any alphabet size.
and you get the theory for any alphabet size.
keiths,
The latest run is certainly an excellent fit to the predictions of the model, owing to the large number of offspring (5000) and the length of the run. That is reassuring, suggesting that the theory is working.
Joe,
It’s gratifying to see a good fit between theory and observation, isn’t it?
I let the run continue overnight. Here are the results after 10 million generations:
I’ll start another run with 50,000 offspring per generation.
petrushka,
Joe’s equations in the OP correspond to the case where the number of survivors is 1, so I don’t even print the predictions when num_survivors > 1.
Why bother? The theoretical results for an infinite number of offspring were very close to those with 5,000 offspring. Why not try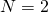 surviving adults instead?
surviving adults instead?
Three in the morning was not exactly the best time for me to mess with the math. I thought I saw that the approximation improves not only with decreases in , but also with increases in
, but also with increases in 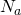 . Perhaps I’m saying no more than that the solution changes little with increases to
. Perhaps I’m saying no more than that the solution changes little with increases to 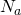 . That’s not a terribly interesting observation. But I’m infinitely more interested in general theoretical results than in whacking my Weasel in public.
. That’s not a terribly interesting observation. But I’m infinitely more interested in general theoretical results than in whacking my Weasel in public.
I’ll check closely, bimeby.
Why not do another OP, and tell us about the Wright-Fisher model for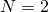 ? I’ve hoped you would get folks to let go of their Weasels, and focus instead on some basics of population genetics. (Fobbing off the monkey/Shakespeare model of cumulative selection as Dawkins’s model of natural selection is a crucial element of ID rhetoric. I’ve long believed that people aid and abet the ID movement when they treat the Weasel as though it were of any importance to the theory of biological evolution. Now I’m in dread of hearing about the Two-Headed Weasel, or some such shit.)
? I’ve hoped you would get folks to let go of their Weasels, and focus instead on some basics of population genetics. (Fobbing off the monkey/Shakespeare model of cumulative selection as Dawkins’s model of natural selection is a crucial element of ID rhetoric. I’ve long believed that people aid and abet the ID movement when they treat the Weasel as though it were of any importance to the theory of biological evolution. Now I’m in dread of hearing about the Two-Headed Weasel, or some such shit.)
Tom English,
With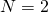 the theory is mush harder because the distribution of matches in the offspring is no longer independent in different positions. The state space is much bigger. Something could probably be done, but not as easily.
the theory is mush harder because the distribution of matches in the offspring is no longer independent in different positions. The state space is much bigger. Something could probably be done, but not as easily.
In the meantime we could look at it empirically with the Wright-Fisher Weasel program,
Tom:
Cheer up, Tom.
The original Weasel demonstrates the power of cumulative selection, and the modified Weasel shows how drift and selection interact. Those are biologically relevant even if the programs themselves aren’t realistic simulations of biological evolution.
The Wright-Fisher model is also biologically relevant though not realistic (infinite offspring, non-overlapping generations), yet you’re comfortable talking about it.
If we had to refrain from discussing any subject that might be misrepresented by someone like Mung, we’d be forced into silence.
The modified Weasel achieves its purpose. It shows that even with a tiny effective population size, drift needn’t dominate selection. It also provides a nice visual demonstration of the population being pushed toward equilibrium, where drift and selection cancel each other out on average, then wandering stochastically above and below that equilibrium point.
keiths:
Joe:
I’m interested in whether the remaining small differences between observed and predicted values are purely stochastic, or whether there is a noticeable systematic component due to the finite population size.
ETA: I’ve also started a run with two survivors to satisfy your curiosity.
I predict that two survivors will make a disproportionate increase in fitness.
Disproportionate to what?
Population genetics theory suggests that two cases that have the same values of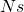 , the same values of
, the same values of 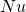 , and are observed on a time scale whose units are 1 per
, and are observed on a time scale whose units are 1 per 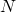 generations should behave approximately the same.
generations should behave approximately the same.
So if we double the number of surviving adults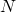 and halve
and halve  and halve
and halve  we should get a process that achieves the same distribution of matches, but takes twice as long to do it.
we should get a process that achieves the same distribution of matches, but takes twice as long to do it.
However this not exact, but is is an asymptotic result. Going from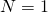 to
to 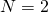 might not show a very close fit. But in general these scaling rules work very well once one gets away from just a few individuals, So the rule would work much better going from
might not show a very close fit. But in general these scaling rules work very well once one gets away from just a few individuals, So the rule would work much better going from 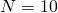 to
to 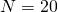 .
.
The mean fitness goes from 16 to 27. See Joe’s post just before this one.
My mean fitness calc doesn’t work with multiple survivors, but the histogram supposedly is unaffected by my add ons.
Did you lower and lower
and lower  as well as increase
as well as increase 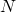 ?
?
And they do treat it that way, even though it is of no importance to the theory of biological evolution. I wish they would stop. 🙂
You have never actually shown this to be the case, you’ve merely asserted it to be so. When you were asked for an empirical test of your claim you declined to produce one.
Whoops.
I’m not confident that my code for calculating the mean fitness is correct when there are more than two survivors.
I get nearly equivalent results with
selection = 5, mutation = .01, survivors = 1
or
selection = 1.55, mutation = .005, survivors = 2
Then this shows the proportional changes not quite giving the same results. I would predict that for larger numbers of survivors the predicted relationship would work better.
Yes. I see a better match between my N=10 and N=20 runs:
Generation: 1318300
Total population size: 5000 Number of survivors per generation: 10
Selection: ON Coefficient: 0.30 Mutation rate: 0.010
Mean Hamming distance: 13.42 Standard deviation: 2.20
Generation: 1599300
Total population size: 5000 Number of survivors per generation: 20
Selection: ON Coefficient: 0.15 Mutation rate: 0.005
Mean Hamming distance: 12.76 Standard deviation: 2.17
And as expected, an even better match between N=100 and N=200:
Generation: 525400
Total population size: 5000 Number of survivors per generation: 100
Selection: ON Coefficient: 0.10 Mutation rate: 0.010
Mean Hamming distance: 11.20 Standard deviation: 2.05
Generation: 526000
Total population size: 5000 Number of survivors per generation: 200
Selection: ON Coefficient: 0.05 Mutation rate: 0.005
Mean Hamming distance: 11.27 Standard deviation: 2.03
keiths,
Would you please check this? I list the number of offspring per parent, so with 1000 offspring per generation for each of 10 parents, there are 10,000 offspring per generation. The mean and standard deviation are for the last 1000 generations.
I’ll run some of yours, a bit later.
************************************************************************
Number of sites L : 10000
Number of possible alleles A : 27
Number of parents : 10
Number of offspring per parent : 1000
Mutation rate u : 0.0001
Selection parameter s : 10000.0
Mean observed num. of fit alleles in parent : 9999.9998
Std. dev. of observed numbers of fit alleles : 0.0141407213395
************************************************************************
CPU times: user 6min 38s, sys: 966 ms, total: 6min 39s
Wall time: 6min 40s