I was short with Joe Felsenstein in the comments section of “Stark Incompetence,” a post in which I address, well, um, the stark incompetence on display in a recent publication of Eric Holloway. I have apologized to Joe, and promised to make amends with a brief post on the topic that he wants to address. Now, the topic is a putative model that Eric introduced in “Mutual Algorithmic Information, Information Non-growth, and Allele Frequency” (or perhaps an improved version of the model). Here is a remark that I addressed to Joe:
Tom English: As you know, if a putative model is logically inconsistent, then it is not a model of anything. I claim that that EricMH’s putative model is logically inconsistent. You had better prove that it is consistent, or turn it into something that you can prove is consistent, before going on to discuss its biological relevance.
I will not have to go far into Eric’s post to identify inconsistencies. After explaining the inconsistencies, which I doubt can be eliminated, I will remark on why the “model” is not worth salvaging. The gist is that Eric’s attempted analysis puts a halting, output-generating simulator of a non-halting, non-output-generating evolutionary process in place of the process itself. An analysis of the simulator would not, in any case, be an analysis of the simuland.
Inconsistency
In the following passage from his post, Eric describes a randomized procedure, dubs it an evolutionary algorithm, and then asserts the existence of a halting program that implements the procedure.
The alleles are 1s and 0s, and the gene G a bitstring of N bits. A gene’s fitness is based on how many 1s it has, so fitness(G) = sum(G). The population consists of a single gene, and evolution proceeds by randomly flipping one bit, and if fitness is improved, it keeps that gene, otherwise it keeps the original. Once fitness(G) = N, the evolutionary algorithm stops and outputs G, which consists of N 1s. The bitstring that is N 1s will be denoted Y. We will denote the evolutionary algorithm E, and it is prefixed on an input bitstring X of length N that will be turned into the bitstring of N 1s, so executing the pair on a universal Turing machine outputs the bitstring of 1s: U(E,X) = Y.
The first thing to note is that the randomized procedure is not an algorithm: it halts with probability less than one. That is, by a simple inductive argument, for all natural numbers ![]() the probability is less than one that the procedure halts after
the probability is less than one that the procedure halts after ![]() or fewer bit flips. Indeed, the probability is greater than zero that the procedure performs
or fewer bit flips. Indeed, the probability is greater than zero that the procedure performs ![]() bit flips without improving fitness. Thus if you were to turn the procedure into an algorithm by stipulating that it performs at most
bit flips without improving fitness. Thus if you were to turn the procedure into an algorithm by stipulating that it performs at most ![]() bit flips, there would be a nonzero probability that gene
bit flips, there would be a nonzero probability that gene ![]() remains equal to input
remains equal to input ![]() when the algorithm halts, no matter how large you make
when the algorithm halts, no matter how large you make ![]() The problem is not the particular randomized procedure that Eric has chosen. Evolutionary procedures do not converge surely to local fitness maxima in discrete spaces (except in trivial cases).
The problem is not the particular randomized procedure that Eric has chosen. Evolutionary procedures do not converge surely to local fitness maxima in discrete spaces (except in trivial cases).
It is gobsmacking for me, though probably not for you (I labored over “Stark Incompetence” in hope that everyone would be as astonished by something coming from Eric as I am by many of his claims), to see Eric flatly assert that the randomized “algorithm” is a program for a deterministic computer. To put it plainly, a universal Turing machine cannot flip bits randomly. Eric’s “model” is inconsistent, and thus is not a model. To get randomness, Eric must
- draw the deterministic bit-flipping program
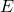 randomly, or
randomly, or - endow the deterministic bit-flipping program
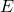 with a pseudo-random number generator, and supply the program with a random seed as input (along with
with a pseudo-random number generator, and supply the program with a random seed as input (along with 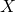 ).
).
However, this will not produce consistency, because Eric has predicated, with the expression ![]() that the program surely halts with an output that is maximal in fitness — no ifs, ands, or buts. His subsequent argument requires it. But his prior specification of the random bit-flipping procedure does not allow it.
that the program surely halts with an output that is maximal in fitness — no ifs, ands, or buts. His subsequent argument requires it. But his prior specification of the random bit-flipping procedure does not allow it.
The map is not the territory
Computer programs used by scientists to model evolutionary processes commonly signal the occurrence of events of interest to the scientists, and halt when the scientists are uninterested in gathering further data. An exceedingly naive response, most prominently on display in the “evolutionary informatics” of Marks, Dembski, and Ewert, is to analyze a program, and claim that the analysis applies to the modeled process — as though the evolutionary process itself signals the occurrence of events and halts.
I am not going to dig into Eric Holloway’s attempt at analysis. You should be able to see for yourself, if you have any business discussing what he has done, that it is literally the program ![]() that enters into the analysis, and not the evolutionary process that is simulated by the program. It ought to be obvious that an evolutionary process does not “know” when fitness is maximized, let alone announce the genome for which fitness is maximal. The part of the program that detects and announces the event in which fitness is maximized, and subsequently halts, is not part of the (simulation of the) evolutionary process. It is a monitor of the simulated process, and can be decoupled from the simulation per se, even though it is usually tightly coupled with the simulation in practice. I advise against struggling to make Eric’s inconsistent “model” into one that is internally consistent, because the result would be a bogus, though consistent, “model” that mistakes the simulation software for the evolutionary process itself.
that enters into the analysis, and not the evolutionary process that is simulated by the program. It ought to be obvious that an evolutionary process does not “know” when fitness is maximized, let alone announce the genome for which fitness is maximal. The part of the program that detects and announces the event in which fitness is maximized, and subsequently halts, is not part of the (simulation of the) evolutionary process. It is a monitor of the simulated process, and can be decoupled from the simulation per se, even though it is usually tightly coupled with the simulation in practice. I advise against struggling to make Eric’s inconsistent “model” into one that is internally consistent, because the result would be a bogus, though consistent, “model” that mistakes the simulation software for the evolutionary process itself.

Unfortunately, that linked quora answer is unclear to me. I refrain from speculating on things I don’t understand, so I’ll stick to AIT and its applicability to evolutionary algorithms. Observers can draw their own conclusions about whether and how said results apply to biological evolution.
Fair enough. QM and Von Neumann entropy is mostly about probabilities…for now…
I’m planning to do an OP with a really good video that applies both Shannon and Von Neumann entropy with the math supporting it.
There are quite a few experiments supporting the notions of quantum mechanics in life systems, such as photosynthesis, bird navigation, sense of smell etc.
The most exciting are enzymes using quantum tunneling in metamorphosis, quantum tunneling in mutations etc. Quantum biology is unavoidable, which means Darwinists will have their hands full with how natural selection selected for particles in superposition, or more than one place at the same time…
Super-natural selection will be doing its magic, again…calculating probabilities…😉
I’d gotten only to the comma when I had, literally, my biggest laugh of the year. A canine pal who is staying with me, while his family travels, was miffed by the racket, and headed for the other end of the house. This really did happen. My reaction was spontaneous, not the result of conscious thought. In fact, it was a couple minutes before I recalled your recent foray into neuroscience, which is the strongest of evidence that you feel no compunction about speculating on things you don’t understand.
The error of yours that I addressed in “Stark Incompetence” was remarkable in that you botched horrendously a simple definition that you not only could have, but should have, drawn from a standard reference. Instead you winged it, without referring to any source at all, and revealed that you really do not “get” mathematical definition in general. (You’ve since said that I’ve poisoned the well, when, clearly, all that I did was to sample the well.) You often step out of your depth, and present yourself as understanding matters that you do not.
If you had understood evolutionary algorithms, then you would not have made the mistake that I identified in the OP. You were not just “sloppy.” You, like many people who have only a passing familiarity with evolutionary computation, thought that genetic algorithms and evolutionary algorithms actually are algorithms.
Your evolutionary “algorithm” is a standard topic in evolutionary computation. If you had known what it is called, and had made the least effort to investigate, then you would not have made the error that you did. Feel free to scramble now, find out what it was that you were addressing, and posture as an expert for the “onlookers.” I will know you for what you are, and continue to laugh heartily at your bungling.
Algorithmic information theory (if you really want to reach those “onlookers” of yours, then you should spell out terms) has provided, since the early 1970s, an approach that is much more appropriate than the one you have taken. So I strongly suspect that your understanding of algorithmic information theory is not nearly as great as you make it out to be.
Bull… malarkey. The question posed to you was the biological relevance of what you had been saying about non-increase (“conservation”) of algorithmic mutual information. You know that an evolutionary process does not register the occurrence of an organism with a maximally fit genome [ETA: and report the genome].
DNA_Jock,
Do you really think that by saying, that some diseases are so debilitating, that children die before they even get to adulthood, that THIS is actually saying something about evolution? That’s comical Jock.
Some people get shot in the streets of LA. Being born in LA could be a deleterious mutation.
Now obviously, according to you creative evolutionists, if a mutation is so deleterious, it will be wiped away from the population. That’s how the genetic code became so strong after all, because natural selection wiped out all the bad codes.
So diseases like this, or mutations that cause one to not want to have sex with members of the opposite sex, they will be wiped out in no time, because, you know, the great power of natural selection and all. So is that what happens Jock?
Now, do you wish to talk about real evolution, or are you content to just claim that if a disease makes you born with no legs, or makes you sterile, then see, we can decide which genes are deleterious to fitness. Oh please.
Some things just don’t change. I’m sure there’s an evolutionary explanation for that.
Merry Christmas Tom and phoodoo.
I just said don’t understand, not understand well. I’ll happily speculate in ways that people can prove wrong. But I don’t know enough about QM to even be wrong.
What do you think is wrong about the neuroscience article? If you have an objection I might be able to test it.
Welcome back Mung!
Merry Christmas, Mung
It’s saying that there is such a thing as deleterious mutations, as these very obvious and rationally undeniable examples demonstrate.
Just to clear up any confusion here I feel like I should state, for your benefit, that being born in LA is not a mutation. In case you really didn’t understand, which I’m genuinely concerned that you didn’t.
Eventually, yes.
Yeah pretty much. Among code-variants, those that made deleterious mutations less likely, eventually took over the populations.
If you’re literally the first person ever born with a mutation that makes you completely refrain from ever having sex and reproducing, then yes you will not pass on that mutation, and so when you die that mutation will go extinct again. Obviously.
You’re the one who suddenly started drooling about “Do you think it would be possible to detect a “deleterious” mutation without any information whatsoever about survival of descendants?”
And the answer is and remains yes, and examples were given. Nobody claims that alone constitutes some sort of final word on evolution by natural selection.
Pretty obviously, yes. It is possible to identify deleterious mutations, some times it is even trivial. And DNA_Jock gave examples.
No phoodoo, I never claimed to be “saying something about evolution”. I was merely demonstrating how easily you and nonlin can be made to write things that are OBVIOUSLY wrong. Thank you for the demonstration.
This is why I enjoy our interactions. You are so delightfully clueless that you offer up great teaching moments. Joe F can provide you with the rigorous math, but simply put, for a recessive lethal mutation, the prevalence of the mutation in the population will reach an equilibrium where the rate of allele removal (due to homozygous individuals dying) will equal the rate of de novo mutations. So they never fully disappear.
Because I am something of an asshole, I deliberately chose highly deleterious recessive alleles of genes on the X chromosome. They get cleared out of the population MUCH faster. And yet we still see them, because (and we sequence the DNA of the parents to confirm this) novel mutations arise every generation. With X-linked inborn errors of metabolism, we get a really strong read on the de novo mutation rate.
These deleterious alleles are continuously getting cleared out, but they are also continuously arising. Hence their prevalence.
I genuinely miss you. Best wishes to you and your family.
Mung,
Mung, Merry Christmas! It’s like the second coming! Have we been good enough?
Welcome home!
Tom English,
I do too.
DNA_Jock,
Well hold on a second there Jock. Your claim was you can know something about a deleterious mutation without knowing anything about any descendants. But of course you do know, they died at childhood and didn’t have any. So that is knowing something about their descendants. You still haven’t passed your own test.
But careful, I can use your definition of deleterious to include all kinds of mutations . Evolution can make a story for anything.
That seems to be equivalent to the old line that “If your parents didn’t have any descendants, then you probably didn’t either”.
Yea, it sure does, and that is the whole problem with your whole fitness concept. You keep claiming its not about counting, but then finally you admit its just counting.
Sometimes you count them when they are young, sometimes you count them when they are old, sometimes you count them before they die, sometimes after they die, sometimes you say they are fit because they are old enough to have offspring, sometimes they are fit because they actually had offspring, and sometimes they are fit because you predict one day they might have offspring. Its all the same thing. I can call any gene that exists fit, by virtue on the fact that it exists, all I have to do is use your counting, and say I counted it.
heck, you can’t even claim being born without reproductive organs is unfit, because all I have to do is say, well, they got old enough to have theoretically had offspring, so since sometimes that is how you count them, that this is fit.
I can also call any gene unfit, because all I have to do is say they have reached puberty and not had any offspring, so , see, I use offspring count as fitness, and they have zero-so unfit.
Jock plays a con game when he says its not about offspring, but he gets caught in his own definition weakness. If a hemophiliac lives to be able to reproduce, they have passed the qualification to be fit. You don’t get to predict they won’t have offspring, just because that’s what happened to others. Its like saying some fat people don’t have offspring, or some college grads don’t have offspring, so I am expecting them not to. You are just picking and choosing your criteria to suit you.
Hey phoodoo, just out of curiosity, do you think you’re the one of all of us who appears to be making the most sense? When you write a post like that, do you truly think you’re somehow showing something wrong with evolutionary biology, and the concept of fitness?
“Anesthetic gases dissolve in hydrophobic pockets by extremely weak quantum mechanical van der
Waals forces known as London dispersion forces (Halsey, 1989). These are instantaneous couplings
of electron clouds between two or more non-polar atoms or moleculesEmutually induced dipoles. In
other words the electron clouds of two (or more) neighboring atoms or molecules (e.g. one from an
amino acid of the protein pocket, and one from the anesthetic molecule) shift instantaneous electron
locations to avoid electron repulsion and maximize attraction between electrons and positively charged
nuclei. The weak binding accounts for easy reversibility—as the anesthetic gas flow is turned off,
concentrations drop in the breathing circuit and blood, anesthetic molecules are gently sucked out of the
pockets and the patient wakes up. But why does such weak binding cause anesthesia?
It turns out that in the absence of anesthetics, i.e. during consciousness, these same weak London forces
occur in the same hydrophobic pockets among electron clouds of amino acids and act to govern normal
protein movement and shape because the various relatively strong bonds in proteins cancel out (Voet &
Voet, 1995). A logical conclusion is that anesthetic-induced London forces perturb normally occurring
London forces in hydrophobic pockets of brain proteins which are necessary for protein conformational
dynamics and consciousness (Hameroff, 1998b). Because the location of an electron is a quantum
process (the location cannot be definite at any time, and in fact is apparently smeared out spatially like a
wave) London forces are quantum mechanical. Thus the underpinnings of neuronal activities are
quantum mechanical interactions. If these interactions are unified in a common wave function then a
quantum field and sequence of quantum events may indeed comprise consciousness.”.
article-template Page 10 of 16
http://www.consciousness.arizona.edu/hameroff/Ham/Ideas/Other_side/other_side.htm
In short, under anesthesia, all brain activity is normal, as indicated by EEG, with the exception of consciousness activity. So, if consciousness involves some kind of a soul, it gets immobilized by anesthetic gases acting on van der Waals quantum mechanical forces…
Mung,
Mung,
Has your life been better or worse without TSZ?
J-Mac,
It needs to be life altering one way or the other?
Who would take it that seriously, it is simply an avenue for hearing other opinions and considering their value or validity one way or the other. For me, its just a way of knowing what evolutionists consider valid evidence for their beliefs. I find most it lacking, so its useful in that respect for me. But hardly life altering.
Its certainly not going to make my life better or worse.
No, but it can be…
It gives me an insight into how bias world view can alter one’s rational reasoning to the point of denying solid evidence against one’s world view…
ETA: It applies to theists, materialists or agnostics
Wrong, phoodoo.
As I explained to nonlin when I originally offered up a nonsense mutation in FVIII, the mutation I described has never been observed; I even noted
What I do have is knowledge of biology, and the importance of the blood-clotting cascade and of lysosomal enzymes. Given that knowledge, I can predict that certain point mutations will be deleterious (whatever your definition) without any knowledge of the historical performance of people carrying that specific mutation. This is what nonlin and you claimed was impossible.
With this out of the way, we can discuss back-mutations.
Blatantly false. You know it HAS BEEN “deleterious” in the past. You know crap about the future. Exactly my point: “fitness” is crap as is “natural selection”, as is “evolution”.
When will you two and anyone else reply with your “fitness function”? But of course YOU CAN’T. Because “fitness” is bogus.
Gee whiz, if you cut someone’s veins, they die. And this makes “evolution” true by way of “natural selection” and “fitness”? Sooo retard.
No “knowledge of the historical performance”?!? Even worse… if possible.
Just pretend all “knowledge of the historical performance” about blood flow has been forgotten somehow and you’re done. Peekaboo level intelligence from above and beyond peekaboo age.
I’ve been pretty clear on this subject. The two mutations that I offered up, one in FVIII and one in I2S, have never been observed before. Yet I can predict that they will be “deleterious”, based on my knowledge of biology.
Not an argument that I have made. Your straw-manning is truly bizarre.
DNA_Jock,
Jock Jock, come on now. It doesn’t matter if you have ever observed it or not, that is not really the point. Of course I can conjure up a mutation which either renders an organism sterile, or dead, and then claim that’s deleterious, because they can’t have descendants. But the only reason it is deleterious is because they can’t have descendants. So you know the status of offspring. They can’t have any.
Because of the definition of fitness, the only thing that can make it deleterious is not being able to have offspring. It doesn’t even matter if you think it is less likely to have offspring, because there is no way to predict how many offspring. They either have or they don’t, that’s it.
So you are trying to prove something that is logically impossible, that something can be fit without offspring, or unfit with offspring. Then you have no fitness meaning.
It’s like claiming something can be blue without necessarily being blue.
There is “inclusive fitness”, and a Wikipedia entry that explains it.
Inclusive fitness counts what an organism contributes to the next generation, even if that organism has no direct offspring. For example, this can be useful with worker bees.
You mean certain mutations in certain organisms and certain conditions. Simple logic shows that the mutations themselves are not intrinsically “deleterious” without a particular scenario that includes the host and environment.
And why are we even talking about your unrealized dream?
And will said organism even be born? And if not, what is “deleterious” about the birth that will never happen?
And what has any of these to do with “fitness”, “natural selection” or “evolution”?
As limited as it is, your knowledge of biology is not what we’re discussing here.
Point is: history informs all your estimates and that’s no Darwinist miracle. It’s just common sense.
Well, I offered up hypothetical mutations, because I predicted (correctly) that if I offered up mutations that had been observed, you would complain that I “had knowledge about the survival of the descendants”. Even so, you still made that complaint.
Now, there’s no way of knowing whether the two nonsense mutations that I offered up have ever occurred; all we can say is that they have never been observed, and entered into a genome database. So, just to satisfy you, I offer up the following:
missense mutations in amino acid Cys84 of I2S.
There are six possible missense mutations to this amino acid. I predict that they are ALL deleterious, and I note that none of them have ever been observed. However, given how many male humans have been born, it’s a safe bet that at least one of these boys carried a Cys84 missense mutation.
Well, my ‘particular’ scenario would be any vertebrate in any environment in which at least some of them could survive to maturity; I don’t find that overly specific.
True, I rely on data and evidence. Your “simple logic” appears to be leading you astray.
What we are discussing is the fact that your claimed tautology
does not exist.
We can look at alleles and, in some cases, predict that they will make their carriers unhealthy (or more healthy…) and we can observe that the frequencies of the alleles change over generations, in a consistent manner. Your supposed circularity is broken by our knowledge of biology.
DNA_Jock,
Fitness=descendants.
You still can’t change that.
All you have to do is experience the evolution at the speed of light, like gibbon genome has, and you can have the fitness of saints…
How else can you explain the accelerated rate of evolutionary chromosomal rearrangement in small apes ranging from 38-52 chromosomes?
Supernatural selection can do it all…
All one requires is faith…😉
https://www.nature.com/articles/nature13679
Thank you Tom. I know you mean it. My mother died. Then my cat died. I don’t know which was worse. 😉
Sorry to hear that…
After those 2 tragedies, then there is CSI…same thing…not even different wrapping…
My sincere condolences Mung, hopefully 2020 has something better in store.
Thank you. It’s amazing the things I’ve accomplished in 2019 as a result of those two sad events. So yes, looking forward to 2020.
I feel for you (believe or not), and hope you’re ok. take care Mung
My thoughts are with you Mung and I hope you’ll be back posting regularly now that the new decade is here 🙂
Mung can’t seem to help himself to be serious even in the moment like this…
Unless of course his mother was an abusive witch…
I am sorry for both of you losses, Mung, I’m glad you were able to find something positive in the year nonetheless.
Abundance of empty words = Fake science.
Said cases are ALWAYS dependent on the environment scenario, hence the alleles themselves (without that particular scenario) are NOT “deleterious” and much less “beneficial”.
Example? With YOUR explanation and link to source.