Mung has drawn our attention to a post by Kirk Durston at ENV. This is my initial reaction to his method to establish the likelihood of generating a protein with AA permease (amino acid membrane transport) capability.
Durston: “Hazen’s equation has two unknowns for protein families: I(Ex) and M(Ex). However, I have published a method to solve for a minimum value of I(Ex) using actual data from the Protein Family database (Pfam),
Translation: I have published a method to solve for a minimum value of I(Ex) among proteins that presently exist.
I downloaded 16,267 sequences from Pfam for the AA permease protein family. After stripping out the duplicates, 11,056 unique sequences for AA Permease remained.
Translation: I took some proteins that actually exist. I implicitly assume that they are a representative, unbiased sample of all the AA permeases that could exist.
the results showed that a minimum of 466 [think he means 433 – that’s the number he plugs in later anyway] bits of functional information are required to code for AA permease.
Translation: the results show that the smallest number of bits in this minuscule and biased sample of the entire space is 433.
Using Hazen’s equation to solve for M(Ex), we find that M(Ex)/N is less than 10^-140 where N = 20^433.
Translation: starting from my extremely tiny sample of protein space, multiplying up any distortions (eg those due to common origin or evolution) and ignoring redundancy, modularity, exaptation, site-specific variations in constraint and the possibility of anything more economically specified than an existing protein, the chance of hitting a 433-bit AA permease by a mechanism not actually known in biology is – ta-dah! – 1 in 10^140.

Durston writes:
Then why does he only use AA permeases instead of all possible protein functions? How many discreet functions actually exist in protein sequence space? How many enzymatic reactions are catalyzed by enzymes, how many exist in addition to the known ones? How many structural proteins exist? How many possible are there?
The Szostak lab found functional proteins (four different ones) in a pool of about 10^11 random sequence proteins (sequences that were 80 amino acids in length). And they only tested for one function (bind ATP), they could have tested for thousands of additional functions, such as chemical catalysts and binding to countless other molcules. They also only tested at one temperarure. Who’s to say what else could be found in that pool of random sequence proteins if they had tested at other temperatures, at different ion/salt concentrations or tested for binding to other molecules or even for the presence of weak chemical catalysts?
It is trivial and easy to find functiona proteins in random sequence space. The Szostak lab proved this experimentally back in the late 90’s and early 2000’s and showed it to be true both for proteins and RNA’s:
https://molbio.mgh.harvard.edu/szostakweb/publications/Szostak_pdfs/Keefe_Szostak_Nature_01.pdf
Functional proteins from a random-sequence library
Anthony D. Keefe & Jack W. Szostak
“Functional primordial proteins presumably originated from random sequences, but it is not known how frequently functional, or even folded, proteins occur in collections of random sequences. Here we have used in vitro selection of messenger RNA displayed proteins, in which each protein is covalently linked through its carboxy terminus to the 39 end of its encoding mRNA1 , to sample a large number of distinct random sequences. Starting from a library of 6 x 10^12 proteins each containing 80 contiguous random amino acids, we selected functional proteins by enriching for those that bind to ATP. This selection yielded four new ATPbinding proteins that appear to be unrelated to each other or to anything found in the current databases of biological proteins. The frequency of occurrence of functional proteins in random sequence libraries appears to be similar to that observed for equivalent RNA libraries2,3.”
80 random amino acids strung together into a protein. Generate 6×10^12 different, random copies, test them all for a single (and extremely biologically important) function: Bind ATP.
Among that starting pool of random proteins 80 amino acids in length, there were four (4) different, unrelated proteins found that could do it. That gives about 1 in every 10^11 proteins capable of binding ATP. Which strongly indicates that as an absolute minimum there is at least one biologically relevant function in every 10^11 80-amino-acid long proteins. (I could stop here already, this is enough to render all of creationism bunk).
Notice how only a single function was tested for in that pool of random proteins. They could have tested millions of different functions (bind millions of other biologically important molecules, tested for catalysis of tens of thousands of different chemical reactions, stabilize phospholipid membranes etc. etc.) – but they only tested for one and found it already to begin with.
AA permeases are part of a larger family of general membrane transport protein. The business part – the ‘permeation’ bit – consists of bundles of amphipathic alpha helix, oriented in such a manner as to generate a channel, with the ‘outer’ sides of the helix having greater affinity for membrane and the inner having greater affinity for water. These are a breeze to generate from random space, and to tune by point mutation and selection.
The ways of generating a single such helix are astro-bleeding-nomical. Having got one, getting a bunch of them is pretty straightforward. Once you have this, you have a generic transporter core which occurs well beyond the AA permeases, in all manner of channels from ion to large molecule.
This is why this ‘bitwise’ analysis is so bloody annoying. Especially from a biophysicist (what is it with biophysicists? Yes, Hunter, I’m looking at you). Treat the protein as if it is only made of primary sequence, with each bit equivalent to on/off binary, except with 20 completely distinct states instead of 2, and every bit along primary sequence having the same significance. It’s a heap o’crap.
The central point of this post, however, is the ridiculous manner used to ‘sample’ protein space. Take tuned, large modern proteins (a sample in itself, of lineages that have not yet gone extinct), many almost certainly related by common descent, and then treat it as an unbiased sample of the entire space(s).
The probability of getting a modern protein in one go could not be less relevant to evolution.
Probability mistakes ‘Darwinists’ make my arse.
Precisely.
In addition to the false “independence” assumption that is baked into any p(a) x p(b) x p(c) calculation, Durston assumes that nascent polypeptides must have the same sequence constraint as that observed in (as you so aptly put it) tuned, large modern proteins.
It is beyond stupid.
I do wonder what would happen if someone applied his technique to, say, a helix-turn-helix motif. There’s an ID research proposal…
Rumraket,
You didn’t mention the size of what the Wizards of ID, ever begging the question with terminology, call the “search space.” I’m at the end of a very long day, but I’ll hazard to say that the number of length-80 sequences over a set of 20 amino acids is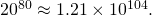 That is, Keefe and Szostak sampled only
That is, Keefe and Szostak sampled only
6000000000000
of the
120892581961462917470617600000000000000000000000000000000000000000000000000000000000000000000000000000000
possible sequences of one particular length, checked for just one particular function, and discovered four proteins that are novel in a very strong sense.
Allan Miller,
I don’t mean to validate the assumption of uniform probability with the calculation I just posted. The point is that the Iddites lose at their own game.
Tom English,
Sure.
I wrote a post on protein space a couple of years back. V J Torley critiqued it at UD, and in my response I detailed some work where just 1.6 million sequences of a ‘pseudo-random’ peptide (based on a 2-state patterning algorithm, to ensure folding) contained 4 sequences that rescued function in E coli knockouts. The total space of the algorithm had 10^56 members IIRC. Assuming these results say what they seem to say, I likened this to using a fragment of a discarded E. coli shell as a scoop, digging into a ball two earth orbits in diameter and finding functional analogues of 4 different randomly chosen modern proteins.
When somebody like Douglas Axe estimates the rarity of functional proteins in amino acid sequence space to be on the order of 1 in every 10^77 proteins, he’s wrong by about SIXTY FIVE ORDERS OF MAGNITUDE. Think about the scale of the error there. Sixty five orders of magnitude.
[TIC]Creationism is removed from reality by about this much:
100.000.000.000.000.000.000.000.000.000.000.000.000.000.000.000.000.000.000.000.000.000
😛
Besides all that, it is known that peptide oligomers as small as three amino acids has potentially biologically relevant functions (the trimer of glycine is a catalyst of peptide bond formation). Will Durston employ his powers of mathematical deduction to estimate the odds of producing a particular three amino-acid peptide?
Should we also now mention that glycine is the most abundant amino acid in biology, the simplest one and easiest to make by abiotic chemistry? Creationism survives partially by a strange but seemingly volitional ignorance of such basic, empirically demonstrated facts.
Allan Miller,
Can you describe this process for me? Does it include helix sequence duplication inside DNA?
colewd,
I did so in part here. This is the fundamental process of generating a single amphipathic helix.
But yes, copying it must involve sequence duplication inside the DNA. Despite stcordova’s protests, everything is made from DNA, and ultimately inherited through that medium, and so any change must take place at DNA level. The article shows several mechanisms by which this can occur.
Getting the duplicated sequence away from its site of origin may involve transposons – selfish genetic elements that copy and paste themselves but may in the process incorporate sequence that is not part of the transposon itself. Equally, viruses can do the same. Or, a DNA fragment may be excised as a loop during homologous repair, which is then re-incorporated elsewhere. Cells seem decidely unhygeinic in their willingness to take up fragments of DNA – that’s how transformation works in bacteria. But it is not necessary for the DNA to be externally sourced. Cells have mechanisms that ensure that subchromosomal fragments do not remain subchromosomal fragments for long – presumably because re-inserting them in the ‘wrong’ place is preferable to losing them, in net terms. You can save whole genes at a cost of some percentage of disruptive incorporation. But sometimes you get a happy result, by actually generating an improvement.
Allan Miller,
You mean with exception of small molecules that are not produced through the TT process?
colewd,
Don’t know what you mean.
Though if you mean water and nutrients … I guess. Somewhat beside the point though.
Allan Miller,
I have done research in the last year on Vitamin d and its biochemical link to cancer. We get vitamin d from the sun, it is processed in the liver and kidney prior to moving from the blood stream to the cells. This modified form is a transcriptional steroid and is a key regulator of the cell cycle. Just an example of a transcriptional molecule that is not DNA based. Just FYI not to make an argument.
colewd,
No we don’t. Vitamin D (other than that ingested in our diet) is generated from cholesterol by enzymes that are produced by transcription and translation from DNA. There is a uv-dependent step, but that’s simply a relatively unusual alternative to activation by ATP, which drives most other energy-consuming reactions. (ATP also, ultimately, derives its energy largely ‘from the sun’. But it too owes its presence to DNA. Ditto cholesterol).
I don’t know what you mean by ‘transcriptional’ there. But … wrong, either way.
Allan Miller,
Saying it is generated in not accurate. The enzymes modify the molecule. What is the origin of the molecule being modified?
colewd,
The cholesterol biosynthesis pathway, another enzyme controlled pathway, of course.
You are being ridiculously pedantic. There is a certain amount of raw material import, of course. But pre-vitamin D is not one of them. You’ll be saying next that DNA is relegated from its central role because it does not create atoms out of nothing.
There is absolutely nothing special about Vitamin D that could not be said about any other biological molecule, beyond the photolysis step. Do you exclude glucose from consideration because it is ‘made using sunlight’ in plants?
colewd,
This incidentally, from the man who says we ‘get Vitamin D from the sun’ … !
Am I in any danger of losing my crown?
Allan Miller,
I think we are agreeing at this point. Thanks for the discussion.
Allan Miller,
Ok…I left out a few critical process steps…my bad 🙂
It’s all in the demand for details!
colewd,
Not sure on what, but cheers!
There are two points to respond to. The first to do with how representative my/nature’s sample size is and the second to do with the Keefe and Szostak experiment.
Regarding representative sample size:
The assumption in Miller’s ‘translations’ is that evolution has not yet had time to provide us with a proper representative sampling of sequence space for the biological proteins. There is a way to test that assumption, which I did several years ago at the beginning of my research. First, it is important to understand that both Hazen’s and Szostak’s and my equations to evaluate I(Ex) are based on Shannon’s equation, with the added co-variable of function. At the heart of the equation is a summation of probabilities of each symbol in the sequence one is measuring. What this means for proteins, is that need to know the probabilities of all 20 amino acids at all the sites in the sequence. So for a 433 residue protein such as AA permease, we need to know a total of 20 x 433 = 8,660 individual probabilities. To estimate these probabilities, we can use multiple sequence alignments provided by Pfam. These Pfam sequences represent sequences that have not been eliminated by natural selection (i.e., are probably functional), and span a wide range of taxa, the wider the better the sampling.
So to test whether we have a representative sample, we need to see how the frequency distribution changes with sample size. We can do this by plotting I(Ex) vs sample size (i.e., number of different sequences). At first the curve drops very steeply as sample size is increased but begins to level out toward a horizontal asymptote. What that means is that the frequency distribution is beginning to stabilize. At that stage, one can double or triple the number of novel sequences and see very little change in I(Ex). I have found that I need at least 500 different sequences before it starts to level out. Preferably, I like to work with at least a few thousand.
If, however, the curve showed little sign of leveling out toward a horizontal asymptote, but kept steadily dropping instead, then that would indicate that our sample size was inadequate. More precisely, it would show that there were still a lot of sequences out there with significantly different frequency distributions than what evolution had discovered. If that were the case, then Miller’s fears would be justified. As it is, for protein families that span a wide range of orders or phyla, evolution has done a very nice job of sampling functional sequence space.
Regarding the Keefe and Szostak experiment:
1. Keefe and Szostak’s results are for a sequence space of 80 amino acids. The average protein consists of 300 amino acids. This requires a vastly larger sequence space 20^220 times as large, on average.
2. Biological proteins that bind to ATP must also be able to release it to complete their function. (See http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2415747/ ) That may substantially raise the bar. They did not test to see if their protein was able to perform that feat in vivo. Their experiment was done in vitro. That is a very large difference. I expect that adding that extra requirement will substantially reduce M(Ex)/N.
3. Binding to ATP is a relatively simple function compared to the much more stringent functions the typical biological protein must perform. One can expect, therefore, that the 1 in 10^11 figure represents an easy target, compared to the more demanding functions the average protein must carry out in living cells.
4. Keefe and Szostaks’ paper was published 15 years ago. Our knowledge of sequence space, and our ability to measure it, has greatly advanced since then. Specifically, our library of functional sequences for thousands of proteins had been increased enormously, to the point where we can use real data to get a much more accurate estimate for M(Ex)/N for different protein families than Keefe and Szostak had available 15 years ago, which is precisely what I am doing. I have recently done six protein domains ranging in size from 33 aa up to111 aa. The actual M(Ex)/N fractions range from 10^-18 for the ankyrin repeat to 10^-100 for the RS7 domain. Keep in mind that these are domains that actually carry out biological functions, unlike the in vitro function in their experiment.
5. Even if we grant their 1 x 10^11 M(Ex)/N for an 80 amino-acid protein, if we wish to construct a 320 amino-acid protein, which is much closer to the size of the average protein, we would need four different 80 aa domains, each with its own area of sequence space distributed within the larger 320 aa sequence space. At best, the probability M(Ex)/N would decrease to (10^-11)^4, or 1 chance in 10^44 in a single trial … but the extreme upper limit for the total number of trials in the history of life is less than 10^43.
6. Finally, it should be pointed out that their experiment is an example of intelligent design in action, requiring very careful, intelligent selection and purification strategies. The experiment shows that if we begin with purely random sequences, we can locate functional sequences if we have a goal in mind (which evolution does not), and we intelligently design a highly controlled series of selection and purifying steps under laboratory conditions. No one disputes that we can, by using our intelligence, locate functional sequences.
Welcome to TSZ, Dr Durston.
Welcome, Kirk.
The sequences were randomly generated. The selection (non-random) found the sequences that bound ATP best. Nobody designed the sequences. This is classic evolutionary theory.
Kirk,
I assume from this you conclude that stochastic processes are very unlikely to be the cause of large scale evolutionary change.
No, this is complete gibberish. There is absolutely no reason to think plotting I(Ex) vs sample size reflects the number of potentially functional sequences in all of sequence space. You simply can’t make that kind of extrapolation.
There’s absolutely no indication that there’s a connection between your calculation and whether there are any unknown but possible functions in unexplored sequence space.
But you’re arguing the opposite, that evolution could not POSSIBLY sample enough of sequence space to discover any functions. Which is the point of this thread, because we agree that evolution can’t have sampled much of the entirety of sequence space, but has nevertheless discovered a lot of functional sequences. This implies the diametrically opposite to what you are trying to conclude: That functional sequences actually exist at frequencies that are many orders of magnitude higher than your estimation.
You have supplied no information that indicates the frequency of functional sequences is any less for 300 amino acid proteins, than for 80 amino acid proteins. So for all we know, the total size could be relatively irrelevant.
At least there is some merit to this question in the sense that it is actually an unknown how it affects the frequency of function in sequence space to add another criterion.
I suspect you could get to “there” from “here” with selection as other experiments by the Szostak lab, with these proteins, indicates.
I don’t see any reason to think the protein wouldn’t be able to bind ATP in vivo, nor that if it couldn’t, the frequency of ATP binding proteins that function in vivo occur at any less a probability.
I don’t agree that it is any more a “relatively simple” function than any other binding activity. It’s a nucleotide binding protein, it is no simpler than proteins that bind to other proteins or to DNA. I suspect these are abundant in sequence space.
It is intersting to note that the Szostak lab did additional selection experiments on these proteins and found that with a few rounds of selection, the protein would be improved to reliably discriminate between ATP, ADP and AMP (it would bind ATP but not ADP and AMP). Combined with the fact that there were four novel ATP binding proteins not related to any sequences known from life, and that high specificity was relatively close in sequences space, this actually indicates that additional functional criteria are relatively close in sequence space and can be easily sampled by mutation and selection in vivo.
This is completely irrelevant information. You are equivocating between the frequency with which functional proteins occur in amino acid sequence space, and the frequency with which specific domains used by extant life occur in sequence space. You are somehow trying to imply the proteins used by extant life are the only possible functional ones. Which is just not true and why I brought that Szostak lab reference, because it directly contradicts that inference by discovering functional proteins completely unrelated to anything known in extant life.
The proteins known from life are constrained in their sampling of the totality of sequence space due, primarily, to common descent from similar ancestral sequences. Evolution works mostly by slow accumulation of change, rather than by totally random sampling. It is remarkable that both are routinely demonstrated to work at discovering novel functions in direct laboratory experiments.
I’m sorry but this is just ridiculous and there is nothing in biochemistry that merits this kind of cumulatively dwindling frequencies of functional proteins as they grow in size.
There is absolutely no reason to think that if the frequency of specific functions exist at a frequency of 1 in 10^11 for 80 aa-proteins, that the frequency of functional proteins would be powers of this. Why do you even believe this?
This one is straight up nonsensical Kirk. It is trivial to draw an analogy from this experiment to a selection pressure that could happen in the wild, such as a foreign antibiotic invading the cytosol of an organism and one of the cytosolic proteins by random having very weak binding affinity of the antibiotic which inhibits it’s antibiotic effect. This would correspond to the random initial sampling part of the experiment (the proteins in the cytosol constitute a sample set, of which those that can bind and thereby inhibit the antibiotic represent “found functions”). The subsequent rounds of selection then just corresponds to generations of bacteria undergoing mutations and eventually culminating in a mutated cytosolic protein strongly inhibiting the antibiotic. Obviously all along the way, those bacteria with cytosolic proteins with stronger inhibitory effects outbreed their competitors carrying weaker versions and so on.
You guys really need to let this particular line of argument go, any thinking person can instantly see through it.
Kirk,
Hi Dr Durston, thanks for responding.
This is not an assumption I knowingly make. Even if given an infinity of time, evolution would not sample sequence space in a representative manner. Indeed, the more time goes on, the less representative the sample is. Evolution, by its very nature, does not give representative samples.
Common Descent first:
At the extreme a single AA permease in LUCA could be inherited by all descendant organisms. So the modern data would reflect the connectivity of a local region of space, that reachable by evolutionary ‘methods’ (point mutation and sub-ORF recombination) from the start sequence, and not the likelihood of the start sequence itself. The sequences passed through on the way or arrived at would be part of the totality of functional sequence, but provide no guide to the largely hidden portion: All Possible Sequences.
I am not proposing that the sampling is that extreme, but the problem should be clear: one would be taking a single instance (the LUCA sequence), copying it with amendment, and claiming that one had thereby sampled the entirety of space. That is a strong sample bias. The real bias may be somewhat less – the modern permeases may have several origins – but one is still multiplying up a bias, where there is any common descent. Eliminating duplicates removes those sequences that have not changed at all, but keeps all those that have changed at one or more sites.
Selection next:
One is frequently pointed to the length of modern proteins. There is no doubt that, in the dataset and alphabet provided, the smallest permease can be reduced to 433 bits. But this cannot be taken to be the minimum length of a functional permease, still less the minimal length attributable to the permease of any ancestor. There does appear to be extensive selection among proteins for functional changes that also increase length. This makes some sense when one considers the various subfunctions associated with a protein. A protein with a certain minimal functionality cannot gain additional functionality using the ‘bits’ it already has – they are fully occupied with current function. So (for example) adding an extra alpha helix to the transmembrane component would inevitably increase the overall length, while simultaneously providing a potential improvement in function. A shorter early permease might not survive a contest with modern ones, but could survive a contest with its own, shorter, ancestor. While this is not a ‘proof’ that selection increases length, it is a possibility that needs to be borne in mind when setting a primordial floor.
Finally, redundancy.
There is extensive substitution capacity within a protein along most of its length. Therefore, a metric involving 20 acids at every site seriously underestimates the density of functional sequences. For example, as far as the transmembrane alpha helixes are concerned, an enormous set of sequences can provide the basic 3.6 turn, end-joined as much as you like. The helixes in modern permeases are amphipathic, which assists in creating a channel, but extensive tuning of hydrophobic moment is possible from a less ‘ideal’ configuration, by serially substituting polar for hydrophobic on the inner surface, and the reverse on the outer. This is not a 20-letter alphabet, at these sites, but 2. Even there, the sites are not digitally specified, but substitution depends upon the overall hydrophobic moment of an entire neighbourhood, which relaxes substitution constraint still further.
It’s nice to have Kirk here. Let’s not get off on the wrong foot.
Here’s what he actually wrote:
That is about as far as you can get from “classic” natural [no intelligence allowed] evolutionary theory. We all ought to be able to agree on at least that much, surely.
You left out the important concept. The sequences were not designed.
The important question is whether sequences have to be designed. Whether they are so distant in sequence space as to be impossible to make without design.
The other critical concept is whether functional sequences are connectable by single mutations.
Kirk did not say the sequences were designed. That’s either putting words in his mouth or creating a straw man. Let’s not abuse Kirk’s visit by making stuff up instead of dealing with what he actually writes.
Once again, here is what he actually wrote with a different selection of the text emphasized:
He didn’t claim the sequences were designed, he claimed they were located.
The claim that was made in the previous thread, which spawned this one, was that the size of sequence space was irrelevant.. Now we are hearing that it is relevant.
Sequences that merely bind ATP in vivo could very well be fatal to the cell.
Actually, no.
petrushka,
or…have we identified the mechanism that is causing large scale evolutionary change?
Mung,
I think you have completely misunderstood Rumraket’s point, if that is how you read it. Evidently, the space must be bigger – much, much bigger – than that currently occupied by real proteins. Outside of that, all that matters is functional density. In fact, for evolution, local functional density – the availability of paths of amendment.
colewd,
No.
Mung,
Leaving evolution to proceed via the things that very well may not.
(I feel I may have to make this point several times more yet) neighbouring sequences do not merely consist of those one or two point mutations away from each other. Segments of sequence can be moved by copy/cut and paste, and reciprocal recombination, both within and between ORFs. This makes a massive difference to the number of paths available. Proteins do not appear to have been assembled by N random picks from a 20 acid library, nor modified solely by bit-position substitution.
It’s silly. The purification steps are done to be able to measure the functionality of the product, not because they’re necessary for the proteins to work. I’ve already explained what is wrong this that line of reasoning (that experiments like these qualify as “intelligent design” rather as an analogue of natural selection) above. There’s nothing about this experiment that implies something like this couldn’t happen in the wild or just in vivo. The only reason anything is being done during in vitro experiments that don’t take place in the wild is so scientists can actually study the properties of the reactions, not to somehow stack the deck.
Irrelevant with respect to what question or claim? It can’t just be blanket irrelevant in any and all circumstances, obviously. If the claim is that evolution has sampled all or a significant fraction of sequence space and found all possible functional sequences, then that claim is just unambigously wrong.
Who made the claim that sequence space was irrelevant and in what context?
Not really. There’s a lot more ATP in your cells than there are individual protein molecules and ATP is continously made in large quantities (millions pr. second). You could have a million copies of a single ATP binding protein and still be very far from exhausting the ATP cache of the cell at any given moment.
Most enzymes actually use ATP and so have pockets that accept ATP where they are phosphorylated. It’s released when it’s converted to ADP or AMP.
Time prevents me from responding to all the comments, so I will limit myself to Allan Miller’s response. My apologies to all the others; I did skim through them at least.
Reading through your notes regarding whether we have a representative sample size, I tend to agree with everything you discussed. The question is whether the evolutionary record in Pfam of successful (i.e., functional) sequences is sufficient to give us a decent estimate as to M(Ex)/N or M(Ex).
A bit about my procedure
I first strip out as many insertions as I can, while recognizing that some of these insertions may actually perform a function in certain taxa. By doing this, however, I increase M(Ex)/N primarily by reducing N. So for the AA permease example, the initial multiple sequence alignment had just under 3,000 columns. After removing insertions, I got it down to 433 columns/sites. My method, however, aims for a maximum value of M(Ex)/N and a minimum value for I(Ex), so the results give me an upper limit for the probability M(Ex)/N, assuming that I have a representative sample of M(Ex).
You raise a good point about an ancestral AA permease being possibly shorter, and performing fewer functions that what we have today. To strip out insertions I use a variable that weeds out all columns that occur less than a certain percentage of times. The assumption is that if the majority of proteins can do without that insertion, then it is not needed to achieve functionality. Of course, it may represent a new function, but I do not know that; I just want to get the length down to ‘average’. Normally, if set the variable at about .15 (weeds out all columns for which less than 15% of the sequences have values), that about cleans out everything that can be cleaned out. Increasing that value does little or nothing to the remaining columns, suggesting that I am getting close to the basic (ancient?) protein length. Not so for AA permease. I had to increase the value to .4 just to get it down to 433 amino acids. This is highly unusual and suggests that the AA permease family contains a lot of functional insertions or that it can tolerate a lot of junk and still be functional. It also leaves room for reducing the size still further, so the ancient AA permease may, indeed, be shorter as you suggest. Further work would be required to see.
Evolutionary sampling of sequence space
For those who have experience writing genetic algorithms, you know that if we just use mutations, we will be stuck sampling only nearby sequence space. Crossover, however, can drop us into a completely different area of sequence space from which a new search area can unfold. Deletions can prune the size of sequence space, giving us simpler solutions. So if evolution can utilize crossover and deletions, it is not stuck sampling ever-widening circles of a local area of sequence space. The question is, how effective has its sampling been?
Why I think my sample size is representative
1. As I mentioned earlier, the I(Ex) vs sample size is approaching a horizontal asymptote. What does this mean? It means that even though evolution continues to provide samples from new areas of sample space through crossover, insertions and deletions, any functional sequences found all fall into a particular frequency distribution as discussed in my earlier post. Conversely, if evolution was still discovering sequences in other areas of sequence space that had different probability distributions for their amino acids or no sequence similarity, the curve would slope down in a more or less straight line with a constant slope. I find that for samples that consist of thousands of unique sequences, the curve is approaching a horizontal asymptote.
2. Protein families have sequence/structure similarity as their defining attribute. Since structure is determined by sequence, proteins within the same family will have a certain level of sequence similarity, which means that evolution need only look for family members in that area of sequence space. With this in mind, it might not be surprising that evolution has had enough time to give a decent mapping of functional sequence space for that particular 3D structure. Other 3D structures in other areas of sequence space may perform the same function, but they would represent a different protein family due to their different 3D structure.
Why I may be underestimating M(Ex)/N: Even though the curve is approaching a horizontal asymptote, it is not perfectly flat, suggesting either there is still the occasional new sequence that does not fit the overall probability distribution of amino acids, or there is a certain amount of erroneous sequences in Pfam.
Why I may be overestimating M(Ex)/N:
– sequences are sorted in Pfam according to similarity by a HMM algorithm. It is very likely that there is a certain percentage of error. This adds noise to the sample and will reduce I(Ex) and increase M(Ex)/N.
– as I mention in my published paper, my method assumes site independence (i.e., there are no higher order dependencies within the sequence). We know that pairwise and higher order dependencies are there. Ignoring them will vastly increase the estimated M(Ex)/N by many orders of magnitude, likely tens of orders of magnitude. Therefore, the probabilities I calculate are optimistic by many orders of magnitude and are much more likely to represent a best-case scenario than a worst case scenario.
Summary: the effect of assuming site independence within the protein structure/sequence is so large that it likely greatly offsets the fact that my I(Ex) vs sample size curve is only approaching horizontal rather than having a zero slope.
Sure. No energy required.
ETA: Dead cells don’t evolve very well.
Then your line of reasoning basically collapses and you simply can’t use these calculations to estimate the density of function in sequence space.
As I wrote above, all you’re doing is estimating the fraction of sequence space that corresponds to some known fold or domain.
Another critically damning issue that undermines the kind of inference you are trying to make, is that function is almost always context specific. To pick something out of a hat, suppose a cyanobacterium finds itself with a snake venom protein. It (in all likelihood) has no use for this protein. In the context of the life of a cyanobacterium, that venom protein is nonfunctional (or possibly even lethal to the cyanobacterium itself). But clearly snake venom proteins aren’t nonfunctional, they’re very useful to snakes. Mutate a gene enough and the resulting protein stops working at what it is doing. Does that mean it is truly without any and all possible function in any and all circumstances? As you say, we just don’t know that.
So it is simply not possible at our current level of knowledge and technology to make even rough estimates at the density of functional proteins in sequence space on the basis of the sequence space occupied by proteins used in extant life.
This brings us back to laboratory experiments as the only alternative, however limited these also are. In this sense it should serve has a hint that a single more or less arbitrarily picked function (bind ATP) scores hits already in the starting population of randomized proteins. You’re getting numbers in your calculations that are tens of orders of magnitude removed from the results of these experiments.
This response is strange because you first suggest ATP binding might very well be lethal in vivo.
Mung,
Leaving evolution to proceed through the ones that don’t die.
Are you proposing a Principle of Universal Lethality of ATP Binding? On what grounds? You can imagine something being problematic, therefore it is problematic?
In fact this study indicates most protein domains originated from a rather small set of primordial peptides that have been recombined to make new domains ever since:
https://www.ncbi.nlm.nih.gov/pubmed/26653858
A vocabulary of ancient peptides at the origin of folded proteins.
Alva V, Söding J, Lupas AN.