“The reason a bike lock works,” explains Meyer, “is that there are vastly more ways of arranging those numeric characters that will keep the lock closed than there are that will open the lock.”
Most bicycle locks have four dials with ten digits. So for a thief to steal the bike, he would have to guess correctly from among 10,000 possible combinations. No easy task.
But what about DNA? Well, in experiments Axe conducted at Cambridge, he found that for a DNA sequence generating a short protein just 150 amino acids in length, for every 1 workable arrangement of amino acids, there are 10 to the 77th possible unworkable amino acid arrangements. Using the bicycle lock analogy, that’s a lock with 77 dials containing 10 digits.
http://www.evolutionnews.org/2015/10/eric_metaxas_on_1100261.html
I believe this is what Mung has been talking about. I asked Mung:
How many goes do you get? How many bacteria in the earth’s soil?
Mung replies:
Not nearly enough.
I feel this is interesting enough for an OP as it seems to finally touch upon what IDers think the designer actually does that can be investigated scientifically.
For example, if we find in a population a protein that is different to the version in an ancestral population but which still works, the by (their) definition, that is prima facie evidence of the designer at work.
Perhaps we can then take the population with the original protein, enclose it in our most sensitive equipment and attempt to detect the designers actions when it “solves the bike lock” and finds the new protein and somehow makes the required adjustment?
If I were an ID supporter these are exactly the sorts of experiments I’d be proposing, and with money on the table (Templeton) I continue to be surprised at the lack of such endeavours. At the very least they can rule out some levels of possible designer interaction at the macroscopic level.
And Mung, I’d be interested in knowing how many would be enough?
Earlier during his direct testimony, Behe had argued that a computer simulation of evolution he performed with Snoke shows that evolution is not likely to produce certain complex biochemical systems. Under cross examination however, Behe was forced to agree that “the number of prokaryotes in 1 ton of soil are 7 orders of magnitude higher than the population [it would take] to produce the disulfide bond” and that “it’s entirely possible that something that couldn’t be produced in the lab in two years… could be produced over three and half billion years.”
http://www.talkorigins.org/faqs/dover/day12am.html

🙂
Allan Miller,
I don’t understand your point here. But can you still dunk a basketball:-)
This point becomes relevant for alternative splicing. Replace the word bytes with exons if you prefer.
This is simply what is required to determine if the hypothesized cause of system x is the real cause. If to demonstrate that ( for argument sake only) that random mutation and natural selection created x then it must also account for the origin of the components. If you want to limit “evolution” to a theory that talks about how post modular exons got organized than that is fair game but ultimately people will ask about the origin of the exon itself.
And has nobody ever thought to ask this? Or have they asked, and the response has been “only magic can meet the statistical requirements”?
Given that colewd has stated that he does “understand that [titan] is made up of alternative spliced exons and tandem repeats”, that does rather deep-six his v^n calculations…
Have the goal-posts moved again?
colewd,
My point is that it is not absolute size that matters for evolution, but the density of function. If an enormous space had 50% hits, we would not be remotely surprised to get a hit every other go. If a tiny corner of the space had 50% hits, but the bulk had zero, we would not be remotely surprised to find we could move around there but not anywhere else.
Well at least that is biological. But modularity is not really about exons. Many proteins don’t have ’em, and there are much smaller motifs. Though, as I see DNA Jock has noted, you have essentially blown your own “Eeets-so-beeeg” contentions out of the water by bringing exons up. If you can shuffle modules wholesale and end up with functional proteins several ways, function is not that hard to get at, is it? Proteins are not nearly so brittle as your digital fantasy would have it.
In the first instance, I am hoping to persuade you to gain a mature understanding of the relationship between digital peptide sequence and function. It seems to be an uphill struggle, despite your claim to be ‘on the fence’.
Forget exons. That is your misunderstanding of what I said. Exons are swapped post-transcriptionally, I am talking of sequence changes within the DNA itself.
People can ask what they like, but ad absurdum every evolutionary explanation ever must be a detailed molecular account starting from the OoL to the feature in question. I’m sure you see how ridiculous that is, but that is what your supposedly lesser claim amounts to.
Mung,
Yes, I’m convinced, no wobbly tables in your house. 🙂
DNA_Jock,
Trying to move those suckers but they are heavy 🙂
v^n only describes the sequential space. I agree with both of you that the relevant issue is how much function is in that space.
With titan the revenant argument is the 20^350 or 4^1100 sequences that had to form to get started. It turns out the titin for both skeletal muscles and heart muscles is quite mutation sensitive. Let say RMNS somehow gets this right. Now we need the code that will create the isoforms based on alternative splicing. We both know we don’t know the sources of that code at this point. So I guess were stuck for now. Where ever the splicing code is, how would you imagine that RMNS created it?
Allan Miller,
I agree here. So far now my reference point is Hunts paper.
I have an open mind here and again believe you have a much better grasp of this then I do.
I do understand that you are talking about protein subunits like alpha helixes but exon formation is part of the evolutionary story per my discussion above. I understand exons are genome components helixes are protein components.
In order to make the theory solid you have to identify how sequences were formed. Between origin of life and us you have at least 32 million possible nucleotides that work together to allow us to function(assuming 10% functionality). Can we start from loosely functioning 15 amino acid poly peptides that somehow organize to greater complexity (Ken Millers explanation to me)? I don’t see this well supported from the evidence we have at this point.
Colewd, my apologies in advance, I know it is rude to comment on someone’s typographical error, but, given your segue below, the typo is too funny on multiple levels not to share:
linked inserted for moar irony.
and as I mentioned previously, titin is made up of 80aa and 100aa repeats…so where does 20^350 come from? And 4^1100? wtf? You do know that the genetic code is redundant, right? and even if it weren’t, 3 x 350 = 1050? But what’s 10^30 between friends?
As you so rightly said, the relevant issue is how much function is in that space.
colewd,
I’m not all that interested in Art Hunt’s paper, inasmuch as it compels me to adopt no particular viewpoint. I am not Art Hunt. My own view on protein space is contained in the OP I pointed you at. Whether that concurs with or contradicts Hunt’s is as likely to be down to your own understanding as to any real point of dispute.
That’s not quite it. Exons are the part of an RNA transcript that is not edited out. Because they have boundaries, they can be added, dropped or shuffled in different isoforms where there is more than one of them. The term ‘exon’ also applies to the underlying DNA sequence – but ultimately they become protein sequence, including alpha helix.
Alpha helixes are, necessarily, part of exons. That is, of course, the DNA sequence of the alpha helix, not the helix itself.
But I am urging you to drop exons because it seems to be the only way you think a piece of protein can be shuffled. It isn’t. Many proteins have no introns (and so can be considered a single exon). Their sequence-space size is just as irrelevant, and they can still be internally shuffled and part-duplicated.
Your fantasy depends upon every amino acid being mutationally and historically independent of every other in every protein. This is not the case. Blocks larger than a single acid but smaller than an exon can be permanently copied and pasted.
So, you have gone from flagella to the entire history of life from OoL to people. Would I be asking too much to ask you to focus?
Take a look at this (be quick – only hosted for 30 days from yesterday).
This is the outcome of a ‘Heliquest’ search I did. It is a search for a particular kind of alpha helix, an amphipathic one. This is one that displays a significant asymmetry between one side and the other of the folded helix. In order to have that asymmetry, there needs to be a significant excess of hydrophobic acids on one side, and polar on the other. These properties tend to cause such a helix to drag its protein to the inside of any nearby lipid membrane. So, if your protein does best attached to a lipid membrane, having one or a few of these helixes somewhere on it will help that to happen.
The naïve view of protein is that it’s all about enzymes, and every bit is essential for catalytic function. But that isn’t the case. Proteins are ion channels, tags, hormones, tensioners, etc, and many parts even of enzymes are not involved in catalysis at all.
Scroll down 1 page and look at the example in the middle (there were 180 hits all told, although many of them were isoforms). You’ve got a sequence IHGLSHSLRQISSQLSSVLS, but looking down the ‘barrel’ of the helix the 3.6 acids per turn mean that every 4th acid is approx. on the same ‘side’. The colour coding indicates hydrophobicity.
The probability of that sequence is nominally 20^20. So, you might think, you have a 1 in 10^26 chance of getting just that 20 bits of that 1 protein. Lotsa zeroes. But look at the colour coding. The yellow residues are Leucine, Isoleucine and Valine. These are virtually the same molecule, as you could determine by checking molecular diagrams. You could replace every such residue by Leucine if you like.
Just making those substitutions, you have a sequence LHGLSHSLRQLSSQLSSLLS
There are just 5 different letters there, so if they were the only 5 amino acids available, you have a random chance of getting the sequence which is roughly the square root of the 20-acid sequence.
Scroll through the list. On my screen, you see a succession of overlaid sequences which all look pretty much alike – hydrophobic residues are anchored to the base of each diagram. The precise letters don’t matter, but there are dozens of functional variations.
We don’t even need a random pick to work first time. Any helix-forming sequence will have a hydrophobic moment (an asymmetry of hydrophobic residues); stochastically, in few of them will it be zero. So if you get a random sequence with at least a hint of asymmetry, and that asymmetry promotes favourable if suboptimal lipid binding, you have an opportunity for selective tuning by point mutation. In next to no time, evolutionarily, preferential substitution of polar acids on one side and hydrophobic on the other will give a nice ‘optimised’ configuration. Optimisation is enhanced by an increase in the alphabet; the latter does not make the job harder.
Having tuned one such helix, propagation of it around other proteins is a breeze.
Allan Miller,
Very cool.
Nicely put. I’m going to keep it in mind to avoid sloppy language when discussing EAs.
It would be interesting to have an actual adult conversation about what is implied by the word search.
My own thought is that once the object of a search is reached, the process halts. That would, in ordinary language, be a defining characteristic of a search.
That would disqualify biology, because genomic change proceeds at a nearly constant rate at all times, regardless of consequences, good or bad. There are variations in rate of change, but not so much in the mode of change.
A search for a missing person often fails to locate the object of the search. Is it not therefore a search?
Jesus, mung, that is just fucking backwards from what I said.
What I said was that a seeming defining characteristic of a search is that the activity of searching stops when the object is reached.
What the fucking fuck does that say about activity that does not reach it’s object?
You are not stupid, and I am not allowed to say what i think about your misreading of this.
I also said that a defining characteristic of biology is that genomic change proceeds at a nearly constant rate, regardless of whether anything useful is found.
To me, that indicates that there is no goal and no halting condition. It does not meet the ordinary definition of search.
It’s a simple question. Does a search have to halt to be a search? You say yes, I say no.
No, that is not what I said.
I most definitely did not say a search has to halt.
What I said was a defining characteristic of a search is a halting condition.
Allan Miller,
Thanks for explanation. My frame of reference is Hunts data and you don’t agree that is a reasonable frame of reference. Your explanation seems credible but I have know idea how to quantify if it makes complex protein and DNA timing control evolution more or less believable.
DNA_Jock,
Thanks for the correction in both cases:-)
Yes, you did.
And there you said it again. What good is a defining characteristic that is not a defining characteristic?
Let me try again:
That is not a defining characteristic of a search and you even admit it isn’t. So why persist in saying that it is?
Sorry if you are having trouble with comprehension.
Searches have a halt on condition statement implied. At least in ordinary usage, which is what I specified.
One could define an exhaustive search of a space. For example, one could use a metal detector to search a field for coins.
In which case, the search ends when the space is covered.
Biology has neither a halt on found condition, nor a bounded space. Genomes change at a steady pace, and nothing at all happens to signify that anything has been found or that the space has been covered.
Allan Miller,
You are solidly demonstrating here you don’t understand the argument. I fully understand there are redundant sequences in proteins.This is the discussion Jock and I were having. This does nothing to solve the sequential space challenge. I give you credit for showing how a simple subsystem could evolve given lots of evolutionary resources but I think you are a long way from establishing a mechanism that can explain how 20000 human proteins evolved. You want to narrow the argument but unfortunately the explanation of the diversity of life is not a narrow one. Allan unless you can connect the dots we do not have a theory and thats where I think we are. Without a mechanism we don’t have a theory. Do you have a quantitative challenge to he Hunt data? Is your defined mechanism still evolution?
Mung,
I click, see that it’s Kirk Durston setting me straight, and immediately guffaw. He’s infamous for his mistakes in probability.
Spot on. And that gets at one of the reasons why Dawkins’s monkey/Shakespeare model of cumulative selection is not a model of biological evolution. In the Wright-Fisher simulation, we saw clear effects of selection coefficient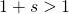 when no individual in the population was close to maximal in fitness.
when no individual in the population was close to maximal in fitness.
colewd,
And you are solidly demonstrating that you don’t, in fact, understand the counter-argument. Redundancy is entirely relevant to the ‘sequential space challenge’. The same stretches of peptide appear all over the protein repertoire, copied from one to the other.
A human gamete is not a pick of 3.2 billion random bits from 4-nucleotide space. So why frame evolution in that way?
I see no point in continuing. It’s not like I win a toaster if I convince you. Anyway, I already have a toaster.
But you may like to wonder why you cannot find a single paper from a molecular biologist in the literature articulating your supposed problem. If you can’t even understand the argument at the local level of an alpha helix (which forms the bulk of many proteins), you are hardly equipped to scale it up. You wish to believe you have found a fatal flaw in evolutionary theory based on sequence space, join the huge club of programmers, comms engineers and physicists who thought the same, deaf to the protests of molecular biologists. May it serve you well.
Mung,
Kirk Durston is full of shit. Next!
Mung
Hahahahaha! I just read what he did! HAHAHAHAHAHAHAHAHAHAHA!!!!
Yes! As in:
Gregory (Scotland Yard detective): “Is there any other point to which you would wish to draw my attention?”
Holmes: “To the curious incident of the dog in the night-time.”
Gregory: “The dog did nothing in the night-time.”
Holmes: “That was the curious incident.”
And, uncoincidentally, full of self-assuredness. I’m disgusted by all who are like him in this regard, irrespective of their particular claims.
Tom English,
Déjà-vu
Allan Miller,
I have repeatably told you I understand this but I still believe you are not going to create these through any mechanism yet on the table. I do think you make the most viable possible counter argument I have heard. Happy to buy you a toaster anyway 🙂 I agree with you that it is time to agree to disagree. Thank you for the dialogue I appreciate it.
Alan Fox,
It was already silly and tiresome back then.
I await the announcement of your Nobel for demonstrating this.
And these are people that Mung rates so highly he sends them money!.
Yep, he rates the DI so highly he sends them money.
I hope you counter by sending the NCSE money.
My bad for thinking Durston was actually “doing science.” LoL
Mung,
There’s a nice new thread on Durston’s post awaiting your defence of his science, should you choose to participate.
colewd,
So you don’t think it possible to generate amphipathic alpha helix from random protein space in the manner I outlined (as a f’rinstance). All amphipathic alpha helix must be designed afresh, or copied and inserted into protein that does not currently have it by this vague commodity ‘intelligence’.
Allan Miller,
I would be interested in the mechanism you think can do this and how it could work. I do think a single component with lose specs and short sequences could fold into a alpha helix with a random process but what is the next step?
colewd,
Short sequences of genome often become duplicated or detached and re-incorporated elsewhere. Too many people are stuck with point mutation as the only method available. If it were all down to point mutation, extensive evolution would indeed be virtually impossible, and bitwise analyses might have some merit.
Allan Miller,
I have read your new op and am brushing up on alpha and beta helixes.:-) Your idea here is solid. Don’t know how close it gets us to a mechanism but worth a shakeout. Will respond on your OP if I have any opinion to add.
If the “search space” is constructed as you and petrishka have been claiming I don’t see why point mutations wouldn’t be enough to get anywhere within it.
There are a couple dozen kinds of mutations. There’s no need to exclude thangis that are known to happen.
But it’s not about reaching a goal. It’s about what has happened an is happening.
Mung,
And the peptide evolution techniques used in drug development rely solely on point mutations. Evolution, OTOH, is like the honey badger…
Mung,
I think there is an excellent chance that you have misconstrued how I think the “search space” is constructed. And, you may not appreciate the enormous rate change, probabilistic effect and increase in dimensionality that recombinational methods cause.
There are vastly more paths available if one is allowed some form of recombination, than if one is restricted to 1-base changes. It is the difference between re-inventing the wheel and re-using it. Protein space is stuffed with re-used wheels.