On Uncommon Descent, poster gpuccio has been discussing “functional information”. Most of gpuccio’s argument is a conventional “islands of function” argument. Not being very knowledgeable about biochemistry, I’ll happily leave that argument to others.
But I have been intrigued by gpuccio’s use of Functional Information, in particular gpuccio’s assertion that if we observe 500 bits of it, that this is a reliable indicator of Design, as here, about at the 11th sentence of point (a):
… the idea is that if we observe any object that exhibits complex functional information (for example, more than 500 bits of functional information ) for an explicitly defined function (whatever it is) we can safely infer design.
I wonder how this general method works. As far as I can see, it doesn’t work. There would be seem to be three possible ways of arguing for it, and in the end; two don’t work and one is just plain silly. Which of these is the basis for gpuccio’s statement? Let’s investigate …
A quick summary
Let me list the three ways, briefly.
(1) The first is the argument using William Dembski’s (2002) Law of Conservation of Complex Specified Information. I have argued (2007) that this is formulated in such a way as to compare apples to oranges, and thus is not able to reject normal evolutionary processes as explanations for the “complex” functional information. In any case, I see little sign that gpuccio is using the LCCSI.
(2) The second is the argument that the functional information indicates that only an extremely small fraction of genotypes have the desired function, and the rest are all alike in totally lacking any of this function. This would prevent natural selection from following any path of increasing fitness to the function, and the rareness of the genotypes that have nonzero function would prevent mutational processes from finding them. This is, as far as I can tell, gpuccio’s islands-of-function argument. If such cases can be found, then explaining them by natural evolutionary processes would indeed be difficult. That is gpuccio’s main argument, and I leave it to others to argue with its application in the cases where gpuccio uses it. I am concerned here, not with the islands-of-function argument itself, but with whether the design inference from 500 bits of functional information is generally valid.
We are asking here whether, in general, observation of more than 500 bits of functional information is “a reliable indicator of design”. And gpuccio’s definition of functional information is not confined to cases of islands of function, but also includes cases where there would be a path to along which function increases. In such cases, seeing 500 bits of functional information, we cannot conclude from this that it is extremely unlikely to have arisen by normal evolutionary processes. So the general rule that gpuccio gives fails, as it is not reliable.
(3) The third possibility is an additional condition that is added to the design inference. It simply declares that unless the set of genotypes is effectively unreachable by normal evolutionary processes, we don’t call the pattern “complex functional information”. It does not simply define “complex functional information” as a case where we can define a level of function that makes probability of the set less than ![]() . That additional condition allows us to safely conclude that normal evolutionary forces can be dismissed — by definition. But it leaves the reader to do the heavy lifting, as the reader has to determine that the set of genotypes has an extremely low probability of being reached. And once they have done that, they will find that the additional step of concluding that the genotypes have “complex functional information” adds nothing to our knowledge. CFI becomes a useless add-on that sounds deep and mysterious but actually tells you nothing except what you already know. So CFI becomes useless. And there seems to be some indication that gpuccio does use this additional condition.
. That additional condition allows us to safely conclude that normal evolutionary forces can be dismissed — by definition. But it leaves the reader to do the heavy lifting, as the reader has to determine that the set of genotypes has an extremely low probability of being reached. And once they have done that, they will find that the additional step of concluding that the genotypes have “complex functional information” adds nothing to our knowledge. CFI becomes a useless add-on that sounds deep and mysterious but actually tells you nothing except what you already know. So CFI becomes useless. And there seems to be some indication that gpuccio does use this additional condition.
Let us go over these three possibilities in some detail. First, what is the connection of gpuccio’s “functional information” to Jack Szostak’s quantity of the same name?
Is gpuccio’s Functional Information the same as Szostak’s Functional Information?
gpuccio acknowledges that gpuccio’s definition of Functional Information is closely connected to Jack Szostak’s definition of it. gpuccio notes here:
Please, not[e] the definition of functional information as:
“the fraction of all possible configurations of the system that possess a degree of function >=
Ex.”which is identical to my definition, in particular my definition of functional information as the
upper tail of the observed function, that was so much criticized by DNA_Jock.
(I have corrected gpuccio’s typo of “not” to “note”, JF)
We shall see later that there may be some ways in which gpuccio’s definition
is modified from Szostak’s. Jack Szostak and his co-authors never attempted any use of his definition to infer Design. Nor did Leslie Orgel, whose Specified Information (in his 1973 book The Origins of Life) preceded Szostak’s. So the part about design inference must come from somewhere else.
gpuccio seems to be making one of three possible arguments;
Possibility #1 That there is some mathematical theorem that proves that ordinary evolutionary processes cannot result in an adaptation that has 500 bits of Functional Information.
Use of such a theorem was attempted by William Dembski, his Law of Conservation of Complex Specified Information, explained in Dembski’s book No Free Lunch: Why Specified Complexity Cannot Be Purchased without Intelligence (2001). But Dembski’s LCCSI theorem did not do what Dembski needed it to do. I have explained why in my own article on Dembski’s arguments (here). Dembski’s LCCSI changed the specification before and after evolutionary processes, and so he was comparing apples to oranges.
In any case, as far as I can see gpuccio has not attempted to derive gpuccio’s argument from Dembski’s, and gpuccio has not directly invoked the LCCSI, or provided a theorem to replace it. gpuccio said in a response to a comment of mine at TSZ,
Look, I will not enter the specifics of your criticism to Dembski. I agre with Dembski in most things, but not in all, and my arguments are however more focused on empirical science and in particular biology.
While thus disclaiming that the argument is Dembski’s, on the other hand gpuccio does associate the argument with Dembski here by saying that
Of course, Dembski, Abel, Durston and many others are the absolute references for any discussion about functional information. I think and hope that my ideas are absolutely derived from theirs. My only purpose is to detail some aspects of the problem.
and by saying elsewhere that
No generation of more than 500 bits has ever been observed to arise in a non design system (as you know, this is the fundamental idea in ID).
That figure being Dembski’s, this leaves it unclear whether gpuccio is or is not basing the argument on Dembski’s. But gpuccio does not directly invoke the LCCSI, or try to come up with some mathematical theorem that replaces it.
So possibility #1 can be safely ruled out.
Possibility #2. That the target region in the computation of Functional Information consists of all of the sequences that have nonzero function, while all other sequences have zero function. As there is no function elsewhere, natural selection for this function then cannot favor sequences closer and closer to the target region.
Such cases are possible, and usually gpuccio is talking about cases like this. But gpuccio does not require them in order to have Functional Information. gpuccio does not rule out that the region could be defined by a high level of function, with lower levels of function in sequences outside of the region, so that there could be paths allowing evolution to reach the target region of sequences.
An example in which gpuccio recognizes that lower levels of function can exist outside the target region is found here, where gpuccio is discussing natural and artificial selection:
Then you can ask: why have I spent a lot of time discussing how NS (and AS) can in some cases add some functional information to a sequence (see my posts #284, #285 and #287)
There is a very good reason for that, IMO.
I am arguing that:
1) It is possible for NS to add some functional information to a sequence, in a few very specific cases, but:
2) Those cases are extremely rare exceptions, with very specific features, and:
3) If we understand well what are the feature that allow, in those exceptional cases, those limited “successes” of NS, we can easily demonstrate that:
4) Because of those same features that allow the intervention of NS, those scenarios can never, never be steps to complex functional information.
Jack Szostak defined functional information by having us define a cutoff level of function to define a set of sequences that had function greater than that, without any condition that the other sequences had zero function. Neither did Durston. And as we’ve seen gpuccio associates his argument with theirs.
So this second possibility could not be the source of gpuccio’s general assertion about 500 bits of functional information being a reliable indicator of design, however much gpuccio concentrates on such cases.
Possibility #3. That there is an additional condition in gpuccio’s Functional Information, one that does not allow us to declare it to be present if there is a way for evolutionary processes to achieve that high a level of function. In short, if we see 500 bits of Szostak’s functional information, and if it can be put into the genome by natural evolutionary processes such as natural selection then for that reason we declare that it is not really Functional Information. If gpuccio is doing this, then gpuccio’s Functional Information is really a very different animal than Szostak’s functional information.
Is gpuccio doing that? gpuccio does associate his argument with William Dembski’s, at least in some of his statements. And William Dembski has defined his Complex Specified Information in this way, adding the condition that it is not really CSI unless it is sufficiently improbable that it be achieved by natural evolutionary forces (see my discussion of this here in the section on “Dembski’s revised CSI argument” that refer to Dembski’s statements here). And Dembski’s added condition renders use of his CSI a useless afterthought to the design inference.
gpuccio does seem to be making a similar condition. Dembski’s added condition comes in via the calculation of the “probability” of each genotype. In Szostak’s definition, the probabilities of sequences are simply their frequencies among all possible sequences, with each being counted equally. In Dembski’s CSI calculation, we are instead supposed to compute the probability of the sequence given all evolutionary processes, including natural selection.
gpuccio has a similar condition in the requirements for concluding that complex
functional information is present: We can see it at step (6) here:
If our conclusion is yes, we must still do one thing. We observe carefully the object and what we know of the system, and we ask if there is any known and credible algorithmic explanation of the sequence in that system. Usually, that is easily done by excluding regularity, which is easily done for functional specification. However, as in the particular case of functional proteins a special algorithm has been proposed, neo darwininism, which is intended to explain non regular functional sequences by a mix of chance and regularity, for this special case we must show that such an explanation is not credible, and that it is not supported by facts. That is a part which I have not yet discussed in detail here. The necessity part of the algorithm (NS) is not analyzed by dFSCI alone, but by other approaches and considerations. dFSCI is essential to evaluate the random part of the algorithm (RV). However, the short conclusion is that neo darwinism is not a known and credible algorithm which can explain the origin of even one protein superfamily. It is neither known nor credible. And I am not aware of any other algorithm ever proposed to explain (without design) the origin of functional, non regular sequences.
In other words, you, the user of the concept, are on your own. You have to rule out that natural selection (and other evolutionary processes) could reach the target sequences. And once you have ruled it out, you have no real need for the declaration that complex functional information is present.
I have gone on long enough. I conclude that the rule that observation of 500 bits of functional information is present allows us to conclude in favor of Design (or at any rate, to rule out normal evolutionary processes as the source of the adaptation) is simply nonexistent. Or if it does exist, it is as a useless add-on to an argument that draws that conclusion for some other reason, leaving the really hard work to the user.
Let’s end by asking gpuccio some questions:
1. Is your “functional information” the same as Szostak’s?
2. Or does it add the requirement that there be no function in sequences that
are outside of the target set?
3. Does it also require us to compute the probability that the sequence arises as a result of normal evolutionary processes?

No it doesn’t. You have to deliberately ignore empirical evidence in order to maintain your view that things always were the way they are now.
Yeah, that’s what phylogenetics is all about.
The core of the spliceosome is made of RNA molecules that are similar to group II self-splicing introns that also exist in prokaryotes. This is evidence right there. The proteins simply assist the splicing mechanism.
There are prokaryotes with self-splicing introns which are similar to the catalytic RNAs of the spliceosome.
A phylogenetic analysis of eukaryotic spliceosomal proteins will show that they’re essentially duplications of each other. That’s evidence of a gradual evolutionary history of the spliceosome right there.
All you can offer here is just some sort of obvious denial, that the evidence somehow doesn’t count. Ahh but phylo-schmylo who cares about the mysterious ways of the designer? That’s just some strange coincidence of design. Or even worse, the appeal to the assumption that some future knowledge will reveal how leaving the appearance of evidence of sequential duplications over geological time is actually a superior method of design. Makes sense? No.
Rumraket,
Lets break this down to workable chunks. Support the claim that PRP8 arose from gene duplication.
You’re not demanding evidence. You started with an implicit claim that it doesn’t exist, apparently based on nothing but the fact that you personally happen to not know about it.
Perhaps worse, when actual evidence is provided we see a move to denial for why the evidence doesn’t count. Mysterious qualifiers are suddenly erected. Oh but the information isn’t “new and original”, or arbitrary limitations are conjured up (oh but the level of function isn’t like the wild-type). Or if another and smaller system can perform the same function, then the system doesn’t exhibit 500 bits but less. And so on and so forth.
Rumraket,
Evidence is fine, hand waving is not. The overall evidence is not supporting the simple to complex hypothesis because as we knock out key elements the cell dies.
The cell is a complex system and eukaryotic cells and prokaryotic cells are very different animals.
Lets look at a simpler problem. Assume first life was a eukaryotic cell. How would you devolve into a prokaryotic cell?
Sure thing. PRP8 nucleotide sequence is homologus to proteins encoded by group II introns. Group II introns contain coding regions called intron-encoded protein (IEP).
See for example
Jessica K Peters and Navtej Toor (2015): Group II intron lariat: Structural insights into the spliceosome. RNA Biol. 2015 Sep; 12(9): 913–917. doi: 10.1080/15476286.2015.1066956
.. and many related articles.
…and is that new and original information or did you just re-use Shakespeare?
Ladies & gentlemen, Bill Cole, gold medalist in the creotard olympics
Yes, and he basically confirms what you accused him of in the first place: gpuccio uses a definition of functional information that excludes information from natural selection.
Mmmmmno, it isn’t like Szostak’s functional information at all.
No it is not. Szostak allows for some non-zero level of function outside of the set that satisfies a specific degree of function. That makes the evolutionary paths (“ladders”) possible that gpuccio denies exist.
Here is the way it was explained in Hazen et al. 2007:
(emphasis mine)
So gpuccio actually admits that he uses a definition of information that excludes the information that evolutionary processes, including natural selection, generate. Here are the additional requirements:
Here gpuccio tries to purge NS from his definition, and thus gives away the game. There is absolutely no way to objectively quantify this *ahem* idiosyncratic type of information. Hence, the 500-bit limit tells us nothing.
Bill,
When you wrote
did you mean that you would add such sequences to the ‘target set’ and then calculate the number of bits?
Thanks.
Your statement is false. 😉
“New” and “original” information is no part of Szostak’s definition of functional information. And in gpuccio’s terminology, “complex” does not mean what we usually think. The organism or molecular structure could be very simple, in colloquial terms, while satisfying the definition of “complex” that gpuccio uses, simply by having the set have probability less than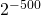 .
.
And since the level of function in nearby sequences just outside the set might be nearly as high as in the set, there would be no implication that you need to get all parts exactly right to have any function.
I re-used Dawkins re-using Shakespeare.
Don’t you think it would be an interesting exercise to calculate the FI for that sequence of letters?
Well, now we know he was wrong.
So once there was a spliceosome, it evolved. I don’t think that is what colewd was referring to. He wants to know where it came from, not what happened to it once it was there. But I think you know that already.
DNA_Jock,
Yes. Any selectable sequence would increase the functional target space and improve the probability of evolving.
To meet gpuccio’s number total sequence space must be 500 bits larger then functional sequence space.
It’s obvious that you don’t have to have everything just right, because there are neutral variants for every working sequence. And from each working variant, other variants. And you can vary the sequence little by little until most of it is different.
Corneel,
If a protein family is novel and does not have a function or sequence similar to any other protein family then what evidence is there for natural selection being involved in the formation of this protein family?
Do we have any evidence of natural selection being involved in the formation of any protein family?
So, what you are suggesting to gpuccio is that it is not the FI of the observed protein that he should measure, but instead he should calculate the FI of the set of sequences with a minimally selectable function from whence the protein could evolve — the FI of the lowlands surrounding the observed peak, as it were.
That’s a very interesting idea, I am curious as to his response.
The greatest quantity of new and original functional information known, by empirical observation, to have arisen without any conscious design intervention is… Damn, I had it right on the tip of my tongue, and then lost it. Someone help me out here.
That portrayal follows from nothing I’ve said. The spliceosome evolved from group II self-splicing introns which exist in prokaryotes (and some eukaryotic organelles). Group II self-splicing introns are not “a spliceosome”. Unless you have your own personal definition of “spliceosome” whereby any putative ancestral entity from which a spliceosomal component of some sort evolved, is also a spliceosome. Is that the case?
Yeah that’s why I explained exactly that.
The real question is: if the members of that protein family happen to evolve by natural selection to a higher degree of function, how do we partition the “new and original” functional information that our Designer injected into the genome from the boring ol’ functional information introduced by random mutation and the selection process?
If you cannot do that, the 500-bit limit is quatsch.
Will you deny the evidence that within any protein family natural selection plays an important role in the diversification of its members?
I thought you would have figured that out by now. It’s 133 bits.
The string has a length of 28 characters and there are 27 characters available for each position. Hence the number of possible configurations is 27^28 ~ 1.2*10^40
“METHINKS IT IS LIKE A WEASEL” is the target string and therefore the only string that achieves this high degree of function (it has a “fitness” of 28 matches).
Hence P = 1 / 1.2*10^40 and the -log2(P) results in a little over 133 bits.
I don’t know whether all that information is “new and original” though.
The absence of evidence for a particular evolutionary hypothesis regarding the ultimate origin of a protein family, is not evidence FOR a design hypothesis.
You understand why that is, right?
Yes. To pick an example, the Thornton lab has elucidated how the GK protein-interaction domain (GK-PID) family evolved from the guanylate kinase family of enzymes sometime around the origin of multicellular animals. They resurrected ancestral states inferred from phylogenetic analysis of the requisite protein families and could show how the GK-PID family originated in a duplication of a guanylate kinase enzyme, and how individual mutations caused a radical functional shift in the protein (it stopped being an enzyme and became a protein-binding protein), and how subsequent mutations significantly enhanced the function.
Read this: Evolution of an ancient protein function involved in organized multicellularity in animals.
DNA_Jock,
His method looks at the range of what has occurred over several hundred million years. If an AA substitution could be tolerated then looking at a variety of animals sequences will show this.
Corneel,
I think your assumption of evolving to a higher degree of function is pretty speculative. This looks like the exception not the rule. Gpuccio’s method explores the range of tolerated mutation over hundreds of millions of years.
Give me some evidence to deny:-)
Rumraket,
If there is no evidence for the claim that weakens the argument for evolution if the claim is relevant. Does it strengthen the alternative hypothesis? Perhaps but it is just a single piece of the puzzle.
That is what his method looks at. As you are aware, this measures permitted variation immediately surrounding the optimum.
But you wrote:
and
That is quite different. Hence my interest in gpuccio’s reaction to your interpretation…
DNA_Jock,
It measures the permitted variation around the function. The “optimum” is your assumption.
That statement is false. It directly follows from what you wrote.
It began with your claim that phylogenetics can tell us how things evolved and continued with your claim about the phylogenetic analysis of the spliceosome in eukaryotes. An analysis that depends on something that already exists (the spliceosome) in eukaryotes.
IOW, it tells us about the evolution of the spliceosome in eukaryotes, but nothing about its purported origin from prokaryotes.
You don’t get that from a phylogenetic analysis of the splicesosome in eukaryotes.
I’ve never had that. Does it taste good?
It hardly matters, since 133 bits is far short of what gpuccio requires for an inference to design.
Thanks for the help with the numbers. So we can’t infer that “METHINkS IT IS LIKE A WEASEL” was designed. As such, Dawkins program failed miserably at showing what he intended for it to show. Go figure.
So what length of text from Shakespear would we need his program to generate in order to hit the 500 bit threshold?
Only with sauce, I believe.
Ouch, that would have upset Shakespeare!
106 characters. “TO BE OR NOT TO BE” is not going to cut it either.
No, they didn’t.
🙂
ok, so say we tried to build a weasel style program with only blind search and random sampling, no selection. The chance it would find a specific string with a length of 106 characters or more? Not in our lifetime? Not in the lifetime of the solar system? Not in the lifetime of the universe? May as well say it is effectively impossible?
Did you read my response?
It would be nice to receive an acknowledgment from you that I was doing no such thing. “Random mutation” and “random variation” is the blind search mechanism. It is only what is produced by the blind search mechanisms that natural selection has to work with. It doesn’t follow at all that natural selection is a blind search mechanism.
You are right. I should have noted that gpuccio’s method looks at permitted variation around an optimum. It may be quite local, and quite temporary. The idea that this represents the global optimum is in fact gpuccio’s assumption, not mine. I’ve tried to point this out to him, without success.
I’ve also tried to point out to him that his “target” should include selectable function that can lead to the observed function, again without success. I applaud your efforts in this regard: maybe he’ll listen to you. Do keep trying.
DNA_Jock,
I think your concept of optimum is ok for single proteins such as enzymes but for multi protein complexes where the protein in question is binding with several proteins to perform a function it is not really coherent. The proteins where gpuccio is finding sequence preservation over time generally fall into this category.
I think the point that Mung is driving home is most relevant. If the total evolutionary resources over the last 4 billion years is 120bits and gpuccio’s design detection bound is 500bits there is plenty of room for yet to be identified islands of function if they exist at all for these protein types.
I see that gpuccio isn’t buying your attempted distinction. You are stuck with the fact that you and he disagree.
Note that he is simply declaring it a “myth” (post #406) that there could be functional sequences outside of his defined target. This is obviously wrong, and there’s plenty of support for Joe F’s position. As you already know.
I encourage you to try to explain this point to him.
DNA_Jock,
From reading more of his posts I think he realizes this is feasible but unlikely to effect the calculation much and certainly within his margin of safety.
Why do you think this a battle worth fighting at this point? I am not sure we have the data to drive this to clarity.
The question he raised that I am pondering is how did these protein parts of the multi protein complexes get into a state where purifying selection can preserve these AA sequences for over a billion years despite lots of DNA mutations.
Corneel, your commentary on gpuccio’s response to me is very astute. Let me
add some comments of my own after your points, and put in bracketed indications of who was speaking in each case.
It is clear that gpuccio’s CFI concept is somehow based on Szostak’s, but as you note it is not the same as it adds further qualifiers.
… [Corneel’s very-apt Hazen quote omitted]
gpuccio says more on that in this later comment. gpuccio seems to assume that a threshold of “function” can be set, such that any level function less than that is too low to be noticed by natural selection. gpuccio uses the very extreme example of an enzyme that makes one molecule of ATP per billion years.
Perhaps people can get gpuccio to explain how gpuccio knows what level of Szostak’s “function” is too low to allow natural selection to take advantage of it. I know enough theoretical population genetics to suspect that the statements gpuccio makes about the ineffectiveness of natural selection are way off base.
The invisible-to-natural selection threshold is certainly a dramatic addition to Szostak’s FI.
It is very simple, Bill.
In order to show that these extra sequences are “unlikely to effect the calculation much and certainly within his margin of safety.” gpuccio needs to “have the data to drive this to [sufficient] clarity.” And he does not.
ID-math consists, fundamentally, of “this calculation is not quite right, but, hey, I reckon it’s close enough, cuz the numbers are reeeally big.”
The issue is that the IDists merely assert that the error in their calculations is immaterial, they never actually demonstrate that the error is immaterial. And yet most of them are engineers — go figure 😮
As a result, ID is vacuous.
Furthermore, you are correct that gpuccio realizes that such sequences are feasible (it is, after all, a necessary consequence of his ‘we could change the threshold’ statements) but he still repeatedly ridicules their existence. Why?
The only motivation I can come up with is that he also realizes that they destroy his calculations, and he is unwilling to publicly admit that.
You should press him on the subject. See how he reacts.
Seems to me that the “new & original” addendum is also damning for his ID “theory”, because you need to specify a function to calculate FI: if precursor sequences had to have a different function, how can you tell how much specified FI has been produced? if you get N bits increase of FI on a transition like that, how much of that was produced for the original function and how much for the final, specified one?
Does that make any sense?
I did miss your response, so thanks, and I will happily retire from the discussion about the meaning of “blind search” as you use it.
I do note that in this comment earlier
you said:
… which does not make it clear that the process of natural selection and mutation is cumulative. The mutation occurs in each generation, starting from where the population is in that generation. That is important.
The cumulativeness is not owing to some mysterious property — it simply is the consequence of the fact that each generation starts from where it is at, rather than back where the starting generation was at.
dazz,
The ‘new & original’ requirement does seem to be formalizing the “first you observe a new and distinct bullet hole, and only then do you apply fresh paint” aspect of the procedure.
So it has that going for it.
DNA_Jock,
There are fudge factors in almost all calculations. We are not in a position to empirically show errors in his calculations until you can demonstrate that a large amount of additional selectable function exists.
The fact that these sequences are mutationally sensitive is very telling. There is so little wiggle room, how would a trial and error process find them?
The funny thing about this and similar OPs is that you can speculate until cows come home and you will never see the end of it because there is no hope that anyone can preform an experiment and say: Aha! Felsenstein was wrong. Or gpuccio was right. Or the other way around…
I’m just not sure why so many people get so excited about speculative “science”… I mean, I like to speculate from time to time when there is no alternative…but I’m not married to my speculations…
Some people here seem to think that speculative “science” is as good as experimental evidence… It is not! I don’t think Joe thinks it is…
Nope. It’s the other way around: we can ignore his calculations until he can demonstrate that they are fit-for-purpose. And, in fact, we can demonstrate that there’s a large amount of additional selectable function. Say 10^93 different ATP-binding proteins, for starters.
Asked and answered.
Have a look at Fig 5 of Hayashi 2006
Not quite true. Of course these guys argue over “How many phenotypes can dance on the head of a pin”, but we still need to positively identify design be it in archaeology, paleontology, cosmology, warfare, etc. And, surprise, surprise, we do… only the method is kind of arbitrary:
Once randomness has been excluded, the system analyzed must be either designed as in “created by an intelligent being”, or a product of necessity as in “dictated by the physical laws”. Currently, a design inference is made when potential human/animal designers can be identified, and a ‘necessity’ inference is made in all other cases, even when there is no known necessity mechanism (no physical laws responsible). This design detection method is circumstantial hence flawed, and may be improved only if a clearer distinction between design and necessity is possible. The DNA-to-Protein algorithm can be written into software that all would recognize as designed when presented under any other form than having been observed in a cell. But when revealed that this code has been discovered in a cell, the allegiance to Darwinism kicks in and those so inclined start claiming that this code is not designed despite not being able to identify any alternative ‘necessity’ scenario.
I say it’s all designed given we know and observe design done by designers but never “design without a designer”. Furthermore, let me reiterate the five full retard claims of “natural selection”
1. “Design by multiple choice” is full retard
2. “Multiple choice from ALL random answers” is full retard
3. “Designing without trying” is full retard
4. “Self design” is full retard
5. “Design by incremental optimization” is full retard