On Uncommon Descent, poster gpuccio has been discussing “functional information”. Most of gpuccio’s argument is a conventional “islands of function” argument. Not being very knowledgeable about biochemistry, I’ll happily leave that argument to others.
But I have been intrigued by gpuccio’s use of Functional Information, in particular gpuccio’s assertion that if we observe 500 bits of it, that this is a reliable indicator of Design, as here, about at the 11th sentence of point (a):
… the idea is that if we observe any object that exhibits complex functional information (for example, more than 500 bits of functional information ) for an explicitly defined function (whatever it is) we can safely infer design.
I wonder how this general method works. As far as I can see, it doesn’t work. There would be seem to be three possible ways of arguing for it, and in the end; two don’t work and one is just plain silly. Which of these is the basis for gpuccio’s statement? Let’s investigate …
A quick summary
Let me list the three ways, briefly.
(1) The first is the argument using William Dembski’s (2002) Law of Conservation of Complex Specified Information. I have argued (2007) that this is formulated in such a way as to compare apples to oranges, and thus is not able to reject normal evolutionary processes as explanations for the “complex” functional information. In any case, I see little sign that gpuccio is using the LCCSI.
(2) The second is the argument that the functional information indicates that only an extremely small fraction of genotypes have the desired function, and the rest are all alike in totally lacking any of this function. This would prevent natural selection from following any path of increasing fitness to the function, and the rareness of the genotypes that have nonzero function would prevent mutational processes from finding them. This is, as far as I can tell, gpuccio’s islands-of-function argument. If such cases can be found, then explaining them by natural evolutionary processes would indeed be difficult. That is gpuccio’s main argument, and I leave it to others to argue with its application in the cases where gpuccio uses it. I am concerned here, not with the islands-of-function argument itself, but with whether the design inference from 500 bits of functional information is generally valid.
We are asking here whether, in general, observation of more than 500 bits of functional information is “a reliable indicator of design”. And gpuccio’s definition of functional information is not confined to cases of islands of function, but also includes cases where there would be a path to along which function increases. In such cases, seeing 500 bits of functional information, we cannot conclude from this that it is extremely unlikely to have arisen by normal evolutionary processes. So the general rule that gpuccio gives fails, as it is not reliable.
(3) The third possibility is an additional condition that is added to the design inference. It simply declares that unless the set of genotypes is effectively unreachable by normal evolutionary processes, we don’t call the pattern “complex functional information”. It does not simply define “complex functional information” as a case where we can define a level of function that makes probability of the set less than ![]() . That additional condition allows us to safely conclude that normal evolutionary forces can be dismissed — by definition. But it leaves the reader to do the heavy lifting, as the reader has to determine that the set of genotypes has an extremely low probability of being reached. And once they have done that, they will find that the additional step of concluding that the genotypes have “complex functional information” adds nothing to our knowledge. CFI becomes a useless add-on that sounds deep and mysterious but actually tells you nothing except what you already know. So CFI becomes useless. And there seems to be some indication that gpuccio does use this additional condition.
. That additional condition allows us to safely conclude that normal evolutionary forces can be dismissed — by definition. But it leaves the reader to do the heavy lifting, as the reader has to determine that the set of genotypes has an extremely low probability of being reached. And once they have done that, they will find that the additional step of concluding that the genotypes have “complex functional information” adds nothing to our knowledge. CFI becomes a useless add-on that sounds deep and mysterious but actually tells you nothing except what you already know. So CFI becomes useless. And there seems to be some indication that gpuccio does use this additional condition.
Let us go over these three possibilities in some detail. First, what is the connection of gpuccio’s “functional information” to Jack Szostak’s quantity of the same name?
Is gpuccio’s Functional Information the same as Szostak’s Functional Information?
gpuccio acknowledges that gpuccio’s definition of Functional Information is closely connected to Jack Szostak’s definition of it. gpuccio notes here:
Please, not[e] the definition of functional information as:
“the fraction of all possible configurations of the system that possess a degree of function >=
Ex.”which is identical to my definition, in particular my definition of functional information as the
upper tail of the observed function, that was so much criticized by DNA_Jock.
(I have corrected gpuccio’s typo of “not” to “note”, JF)
We shall see later that there may be some ways in which gpuccio’s definition
is modified from Szostak’s. Jack Szostak and his co-authors never attempted any use of his definition to infer Design. Nor did Leslie Orgel, whose Specified Information (in his 1973 book The Origins of Life) preceded Szostak’s. So the part about design inference must come from somewhere else.
gpuccio seems to be making one of three possible arguments;
Possibility #1 That there is some mathematical theorem that proves that ordinary evolutionary processes cannot result in an adaptation that has 500 bits of Functional Information.
Use of such a theorem was attempted by William Dembski, his Law of Conservation of Complex Specified Information, explained in Dembski’s book No Free Lunch: Why Specified Complexity Cannot Be Purchased without Intelligence (2001). But Dembski’s LCCSI theorem did not do what Dembski needed it to do. I have explained why in my own article on Dembski’s arguments (here). Dembski’s LCCSI changed the specification before and after evolutionary processes, and so he was comparing apples to oranges.
In any case, as far as I can see gpuccio has not attempted to derive gpuccio’s argument from Dembski’s, and gpuccio has not directly invoked the LCCSI, or provided a theorem to replace it. gpuccio said in a response to a comment of mine at TSZ,
Look, I will not enter the specifics of your criticism to Dembski. I agre with Dembski in most things, but not in all, and my arguments are however more focused on empirical science and in particular biology.
While thus disclaiming that the argument is Dembski’s, on the other hand gpuccio does associate the argument with Dembski here by saying that
Of course, Dembski, Abel, Durston and many others are the absolute references for any discussion about functional information. I think and hope that my ideas are absolutely derived from theirs. My only purpose is to detail some aspects of the problem.
and by saying elsewhere that
No generation of more than 500 bits has ever been observed to arise in a non design system (as you know, this is the fundamental idea in ID).
That figure being Dembski’s, this leaves it unclear whether gpuccio is or is not basing the argument on Dembski’s. But gpuccio does not directly invoke the LCCSI, or try to come up with some mathematical theorem that replaces it.
So possibility #1 can be safely ruled out.
Possibility #2. That the target region in the computation of Functional Information consists of all of the sequences that have nonzero function, while all other sequences have zero function. As there is no function elsewhere, natural selection for this function then cannot favor sequences closer and closer to the target region.
Such cases are possible, and usually gpuccio is talking about cases like this. But gpuccio does not require them in order to have Functional Information. gpuccio does not rule out that the region could be defined by a high level of function, with lower levels of function in sequences outside of the region, so that there could be paths allowing evolution to reach the target region of sequences.
An example in which gpuccio recognizes that lower levels of function can exist outside the target region is found here, where gpuccio is discussing natural and artificial selection:
Then you can ask: why have I spent a lot of time discussing how NS (and AS) can in some cases add some functional information to a sequence (see my posts #284, #285 and #287)
There is a very good reason for that, IMO.
I am arguing that:
1) It is possible for NS to add some functional information to a sequence, in a few very specific cases, but:
2) Those cases are extremely rare exceptions, with very specific features, and:
3) If we understand well what are the feature that allow, in those exceptional cases, those limited “successes” of NS, we can easily demonstrate that:
4) Because of those same features that allow the intervention of NS, those scenarios can never, never be steps to complex functional information.
Jack Szostak defined functional information by having us define a cutoff level of function to define a set of sequences that had function greater than that, without any condition that the other sequences had zero function. Neither did Durston. And as we’ve seen gpuccio associates his argument with theirs.
So this second possibility could not be the source of gpuccio’s general assertion about 500 bits of functional information being a reliable indicator of design, however much gpuccio concentrates on such cases.
Possibility #3. That there is an additional condition in gpuccio’s Functional Information, one that does not allow us to declare it to be present if there is a way for evolutionary processes to achieve that high a level of function. In short, if we see 500 bits of Szostak’s functional information, and if it can be put into the genome by natural evolutionary processes such as natural selection then for that reason we declare that it is not really Functional Information. If gpuccio is doing this, then gpuccio’s Functional Information is really a very different animal than Szostak’s functional information.
Is gpuccio doing that? gpuccio does associate his argument with William Dembski’s, at least in some of his statements. And William Dembski has defined his Complex Specified Information in this way, adding the condition that it is not really CSI unless it is sufficiently improbable that it be achieved by natural evolutionary forces (see my discussion of this here in the section on “Dembski’s revised CSI argument” that refer to Dembski’s statements here). And Dembski’s added condition renders use of his CSI a useless afterthought to the design inference.
gpuccio does seem to be making a similar condition. Dembski’s added condition comes in via the calculation of the “probability” of each genotype. In Szostak’s definition, the probabilities of sequences are simply their frequencies among all possible sequences, with each being counted equally. In Dembski’s CSI calculation, we are instead supposed to compute the probability of the sequence given all evolutionary processes, including natural selection.
gpuccio has a similar condition in the requirements for concluding that complex
functional information is present: We can see it at step (6) here:
If our conclusion is yes, we must still do one thing. We observe carefully the object and what we know of the system, and we ask if there is any known and credible algorithmic explanation of the sequence in that system. Usually, that is easily done by excluding regularity, which is easily done for functional specification. However, as in the particular case of functional proteins a special algorithm has been proposed, neo darwininism, which is intended to explain non regular functional sequences by a mix of chance and regularity, for this special case we must show that such an explanation is not credible, and that it is not supported by facts. That is a part which I have not yet discussed in detail here. The necessity part of the algorithm (NS) is not analyzed by dFSCI alone, but by other approaches and considerations. dFSCI is essential to evaluate the random part of the algorithm (RV). However, the short conclusion is that neo darwinism is not a known and credible algorithm which can explain the origin of even one protein superfamily. It is neither known nor credible. And I am not aware of any other algorithm ever proposed to explain (without design) the origin of functional, non regular sequences.
In other words, you, the user of the concept, are on your own. You have to rule out that natural selection (and other evolutionary processes) could reach the target sequences. And once you have ruled it out, you have no real need for the declaration that complex functional information is present.
I have gone on long enough. I conclude that the rule that observation of 500 bits of functional information is present allows us to conclude in favor of Design (or at any rate, to rule out normal evolutionary processes as the source of the adaptation) is simply nonexistent. Or if it does exist, it is as a useless add-on to an argument that draws that conclusion for some other reason, leaving the really hard work to the user.
Let’s end by asking gpuccio some questions:
1. Is your “functional information” the same as Szostak’s?
2. Or does it add the requirement that there be no function in sequences that
are outside of the target set?
3. Does it also require us to compute the probability that the sequence arises as a result of normal evolutionary processes?

I am assuming that you mean the individual sequences of a larger configuration (say a subunit of a protein complex). Then the answer is: No, we can’t. Because the individual sequences are complex systems in their own right. Hazen et al. explicitely mention RNA polymers as an example:
Of course, its function is different from that of the larger configuration.
Well, I agree with what you wrote here, but I still don’t understand how this prevents mutations from changing the FI in a population. A mutation that changes a sequence to another sequence with increased function will prompt us to change the threshold for that sequence. As it has a higher function, it belongs to a less common set and will be associated with a higher FI. The FI associated with the non-mutated sequences remains unchanged, of course. The average FI for the population of sequences will have increased.
So where do you see a problem?
So you do the exact mistake DNA_Jock says, you assume an unconserved position in a protein can be anything, and that any combination of unconserved positions are viable. Which is strangely enough an assumption that undermines your own conclusion.
Then you also assume that there is zero pleiotropy, so that none of the conserved positions are allowed to change under any circumstances, as in you have simply assumed that compensatory mutations are impossible for conserved residues.
Both these assumptions are completely unjustified, and for any other protein known where experiments have been done, they have been proven to be false. It is NOT the case that all unconserved positions can change to any amino acid, nor can all combinations of amino acids at unconserved positions result in a functional protein. And for the conserved residues, for all proteins where such experiments have been conducted, compensatory epistasis is rampant.
Importantly, neither assumption you make can be used to show that the protein in question can’t evolve. And assumptions don’t give you information about the shape of the fitness landscape for the protein’s function.
Why? Try to explain why you think this. You are not explaining how you arrive at this conclusion, all you are doing is taking the known data, and then stating a conclusion that you have not made it clear must follow from that data.
What I’m missing from you is a chain of reasoning where you show that the conclusion [we are looking at a small pimple surrounded by a very large ocean] follows from the premise [we see different family members clustered at different locations in sequence space, and the sequences represented by each cluster is conserved at X% over long timescales].
Btw, X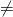 90%. I showed you an alignment for F-type ATP synthase subunit beta earlier in this thread, and for about 100 species it came down to about 15% conservation. And it’s very likely to be even lower given that there were over 5000 more beta subunit sequences on uniprot.
90%. I showed you an alignment for F-type ATP synthase subunit beta earlier in this thread, and for about 100 species it came down to about 15% conservation. And it’s very likely to be even lower given that there were over 5000 more beta subunit sequences on uniprot.
This is very helpful. When you write with clarity, your errors become clear.
You have mis-understood Hazen et al, 2007. They talk of the FI of a ‘system’, where a ‘system’ consists of a set of all possible ‘configurations’ and a definition of function. They also talk of ‘degree of function’.
I have been using the term ‘sequence space’ to refer to the set of all possible configurations, and ‘sequence’, ‘element’ or ‘member’ to refer to individual configurations. Likewise, I have used the term ‘level of activity’ where Hazen uses ‘degree of function’. I hope this has not confused you.
Your error comes from a misreading of Hazen. When the authors talk of the FI of a ‘system’ they are describing a two-dimensional plot, that describes the relationship between Ex and I(Ex) for configuations within that system. There is no FI value associated with any ‘system’. The FI of the system is not 7 bits. It is 7 bits for 00000000 and for 11111111, and zero for all other configurations (for your dichomotous definition of ‘function’; in my system some other configurations have lower non-zero FI).
Here is Hazen on the subject:
I hope you will notice how they talk of the maximum and minimum FI’s of a system, and how, within a system, FI varies with E. The other half of your misunderstanding is your including a threshold value in your definition of a ‘system’. When you calculate a value for what you call the FI of the system, you are actually calculating the FI for configurations that have your ‘minimum threshold’.
The FI of a system is a mathematical function that maps degree of function(activity) to FI values within the context of that system. It takes values only for specific degrees of function.
E2fixtags
You’re like a spectator at some sporting event who is cheering for the home team without knowing anything else about what is going on. Rah! Rah! Rah!
But n is supposed to be the number of sequences at or above the threshold. How is it that before you decided to calculate the FI for each individual sequence that n was equal to 2 and after you decided to calculate the FI for each individual sequence n became 256. I don’t follow that.
I guess what is throwing me off is that I would expect you to use the exact same formula to calculate the FI of 11111111 (an individual sequence) that you use to calculate the FI of 00000000 (an individual sequence) and that you use to calculate the FI of 00011000 (an individual sequence). But I don’t see you doing that.
Could you clarify how you arrive at 256/256? Thanks.
The answer to your question to Joe lies in the sentence immediately preceding the sentence that you quoted:
It is also to be found in the Hazen quote I provided, where they calculate I(Emin) = 0
When you behave in this manner, you appear to be trolling.
Rumraket,
The assumptions are not completely unjustified. Your statement that experiments have proven them to be false is nonsense. An experiment on protein A proves nothing about protein B. This is an indirect measurement and makes assumptions and approximations but is probably conservative.
When you try to shift a burden to the point your opponent has to prove a negative it makes your position look very weak.
Design has better explanatory power as we know that conscious intelligence can generate large amounts of FI and can explain how a protein family can have a member with a very different sequence that finds itself in a preserved position.
Sure it does. For dozens of proteins I have profiled.
Corneel,
Observing jumps in information is not a claim about a design strategy.
I just marvel how you can repeat the mantra “”Design” has better explanatory power” when “Design” explains nothing. It is an empty set. Zero predictive power. No hypothesis that is testable. Come on, Bill. Tell me how a “Design” explanation works. Be the first person ever to do so! 😉
Alan Fox,
It’s been quite a few years. Can you cite any experiments that estimate the probability of binding multiple molecules? How about binding multiple proteins?
Arguments that rely on absence of evidence are not terribly convincing.
This is where your claim falls apart. Szostak’s result shows an island of a 3 inch diameter in the Atlantic ocean. Now how does that help the evolutionary inference?
Alan Fox,
This is as weak of an argument as trying to force your opponent to prove a negative. Design explains FI. Are you trying to claim that FI is nothing?
Blind unguided processes have no predictive power. Design predicts design rules which cancer research depends on.
We have unlimited observations of design creating FI. You and I having this exchange is living proof.
Given that the last time you accused me of trolling I wasn’t trolling, that means little to me. You’ll need to develop a better track record.
In the quote you provided when they calculate I(Emin) = 0 “all possible states” appear in the numerator, such that N = a^L, and this is what Joe has done in his math where he has 256/256. Yet in the example we are discussing that is not the case. Two states DO NOT have “no function.” So I think it’s perfectly reasonable of me to ask for clarification.
Did he mean 254/254?
So let’s cut to the chase. Joe has changed the level of function that he is using to calculate FI. Every possible sequence now has FI=0, and not just “the rest” of them.
Right or wrong?
ETA: So Joe changed the level of function he was using in order to calculate FI, and that’s how he was able to arrive at FI=0. Are we agreed?
Mung,
From my perspective Jock (a competent poster) was trolling in this instant. I do look forward to your continued exchange with Joe on this subject.
I was not accusing you of trolling, Mung, I was merely offering a piece of advice about how you are coming across and therefore how other commenters (or even moderators 😮 ) might view your contributions to this thread.
My apologies. Mung.
I had assumed that you understood the meaning of the symbol ≥
it means “greater than or equal to”.
If the activity level we are considering is the lowest activity level, then ALL POSSIBLE CONFIGURATIONS will have an activity level that is equal to, or greater than, the lowest activity level.
Now I understand why you accused me of a non-sequitur early.
Re-read the Hazen quote, and what Joe wrote.
When he wrote 254 + 2 = 256, he meant 254 (equal) + 2 (greater than) = 256
Alan is clearly saying that Design is not an explanation, not that FI is nothing. Design is not an explanation, it’s a god-of-the-gaps argument whereby you claim that no natural process can produce something that you find a plenty, oddly enough, in nature, yet you claim to be found only in designed objects, therefore god-did-it.
Of course that “blind unguided processes” would have no explanatory power. However, natural processes, with their mixture of random and non-random phenomena, have enormous explanatory power. So much so that we know, in principle, that they work to explain what we see, we only need to figure out exactly which processes and the particular histories followed towards what we see.
Really? Can you point to the ones that didn’t depend on what you claim to be the products of design in the first place? Design that didn’t need energy flow? Design that didn’t need beings that are composed of the very things you claim to be the product of design? Etc? You’re putting the cart before the horse Bill. You’re leaving aside a very deep philosophical problem, and you hold to a very misunderstood vision of nature. Nature is not mere randomness Bill. It has random events, but it also has phenomena with direction. Think of gravitation, think of electromagnetism, think of light propagation, think of energy flow. All of those have directions. Designers depend on those properties of nature. Therefore, it’s nature first, designers much later, as a very tiny part of it.
Simple logic Bill, since we depend so deeply and intrinsically on nature, at such as enormous scale, we are nature’s tiny children. There’s no way around this conundrum for the defenders of the design bullshit.
This is false.
Yes. I get that. And I agree with that. And in that case ALL POSSIBLE CONFIGURATIONS have FI=0. Say it is not so. N = N.
Joe was saying that the rest of the sequences,the remaining 254 of them, (i.e., other than the original two) have FI=0, but what he actually showed was that ALL the sequences have FI=0.
So he was wrong.
To compute the FI for the two, we use the formula …
To compute the FI for the rest …
What, in that case we use a different formula? Why?
And then when we compute the FI for the rest we find we are really computing FI for them all, which is what I have been saying all along.
He moved the threshold. So different answer. But that answer applies to all the sequences, not just to some of them, which was the clear implication of his words if not his math. Which is why I was asking him to clarify.
You clearly don’t find any issue wither the little sleight of hand he pulled, but I do. He was asked to compute the FI for each individual sequence given the original minimum degree of function (threshold), not to do it using a different minimum degree of function (threshold).
At least I seem to be making progress with DNA_Jock. I don’t think he was trolling. He was looking at the math, I was looking at the words and the math.
If Joe had just come out and said, well, to get FI=0 for the rest of the sequences we have to change our minimum so that all sequences are greater than or equal to that minimum…but in that case all 256 of them get FI=0 and not just 254 … so forget about the number I originally came up with those two…
But that would have given away the game.
Poor dazz though. But at least he can still be a cheerleader.
Entropy,
Design is the best explanation for FI and that is what we are observing.
Your philosophical argument is interesting despite the fact it contains materialist assumptions, but non the less inductive reasoning is pointing us to design.
Mung,
If he would respond without his little put downs I would agree. There was no good reason in this instance to not let Joe answer your comment.
Mung,
Makes sense to me. I am grateful for both your, Joe’s and Jocks comments as they stimulate thinking and as such better understanding.
Dazz has made contributions in the past but it has been a while since he has made an argument.
No Mung,
Joe has been correct and you have been incorrect all along.
Here’s a tip: lose your “minimum threshold” concept. The threshold is the activity level of the configuration that you are considering. The threshold is not an attribute of the system.
FI is a property of this level of activity (‘degree of function’ in Hazen) calculated within the context of a ‘system’. FI is not a property of the ‘system’.
When we consider the activity level zero (or whatever the minimum level of activity is within a given system; in my example, the lowest “frequency of the most prevalent character” attainable is four) we calculate that the FI for any sequences having this activity level is zero. Performing this calculation does NOT magically change the FI of sequences that have a different activity level.
Re-read the Hazen quote. They make it really clear that I(Emin) = 0
without qualification, for any system you could imagine.
I think this is nitpicking, if not outright mistaken. It certainly fails to describe any real error of understanding on my part.
You already grant that they speak of the functional information of a system. They also say it [the functional information of a system] can be measured. Presumably this “measurement” produces a value of FI for the system as a fraction of all possible configurations.
They flat out say in their abstract that FI is a measure of system complexity. Perhaps you should take that to heart.
Also from the abstract:
That probability would be a value. If not why not?
In that context it makes no sense to speak of the FI of individual sequences.
Further:
I don’t just make this shit up, I get it from their paper.
So what error?
Accusations that other posters are lying are against the rules. Sleight of hand: “the use of dexterity or cunning, especially so as to deceive”
Yes they are, completely. I note that you don’t go on to detail what is supposed to justify them, rather you just turn to complainabout drawing inferences about protein A from experiments on proteins other than A.
No they’re not nonsense, they’re demonstrable facts. Exactly as I said, for every protein where an experiment has been done that yields data with implications for your assumptions, those assumptions were false for the proteins tested. This isn’t some sort of fluke, it’s true every time protein evolution is tested for some protein.
Uhh that’s just false. There are many aspects of proteins that generalize. While proteins obviously aren’t identical and can vary wildly in their structures and functions, they still have enough commonalities that general rules for the properties of protein biophysics can be elucidated and they apply to all proteins. Fat-soluble proteins all have commonalities, water-soluble proteins all have commonalities, halophilic proteins have commonalities, thermostable proteins have commonalities, and so on and so forth.
This same thing is true for things like how proteins retain their folds and functions despite mutations, and how previously neutral mutations can open up other positions in the protein to changes they would not otherwise have been able to. I have already cited multiple papers in this thread that detail experiments that show this phenomenon. Read everything from the Thornton lab. Twice. Read the references too.
There you go making shit up again.
But you’re the one claiming to offer a method that does exactly that. It’s not my problem when you are the one demonstrably taking on that burden yourself. You claim that when we can calculate 500 bits or more of FI, then we can infer design because evolution couldn’t have done the job. That is you making the claim that evolution couldn’t have done the job.
You’re not JUST saying “Design could have done it”. You’re saying “Because there is 500 bits or more, evolution couldn’t have done it so design must have”.
You can’t complain when I then point out that the evidence before us doesn’t support your claims.
That’s hilariously inept. ID has exactly the opposite of explanatory power because it doesn’t make any predictions. It doesn’t work to render a particular set of data or observations less suprising, nor does it account for any particular patterns in the data over any others.
By the completely vacuous statement that “that’s what the designer wanted”. But you could say that about anything. Suppose we didn’t find that? Well, that’s what the designer wanted. Total lack of conservation? That’s what the designer wanted. Clusters of conserved sequences? That’s what the designer wanted. Scattered dissimilar sequences? That’s what the designer wanted.
That’s not explanatory power. There is no explanation going on.
An explanation is something that makes you understand how or why something is the way it is. Why are there these long lines of fine dust and dirt next along the sidewalk on the street? Well you see it’s been raining these last few weeks, and the rain drags a lot of dirt with it as it flows over the tiles and asphalt, and eventually when it stops raining there is no more water to carry this sediment to the storm drain, so it settles in pools and narrowing streams that eventually disappear leaving these spots and lines of dirt.
That’s an explanation. Design can in principle offer explanations. But they don’t look like this: “it was designed”. A design-explanation has to actually explain something.
A design explanation for something would involve explaining what the designer actually did. What thoughts it had, how it figured out how to do what it did, how it manufactered it’s components, where it got it’s materials. How did you get that stone to have that shape and color? Well first I hacked on it like this, then polished it like this, finally I washed it with this. How did you design and build that house? Well you see I’ve been here before and I know the sun comes up from over there and I wanted my terrasse facing the sunset, but I also needed a foundation because the ground around here can loosen due to seasonal climate bla bla. I bought some stones from a quarry and cut down those trees and stripped them of… etc. etc.
That’s an explanation. It explains what the hell it is we see. “500 bits, which a conscious mind can make” isn’t an explanation.
First of all, you haven’t “profiled” shit. To profile a protein you have to do lab work.
You have compared sequences by copy-pasting into a browser window, to see how many identical residues there were in a pairwise sequence alignment. Second, that actually ignores all the other species where this sequence could have potentially mutated.
You do multiple sequence alignments to get an indication of conservation across the history and diversity of life. You don’t get that from a pairwise alignment comparing just two species. If you just compare (say) human and E coli you might mistakenly think that none of the identical residues can change. Yet that is exactly what you see happening when you sample more and more species to include in your alignment. Now what you will also invariably discover when you do that, is that epistasis is rampant. It is almost always the case that for a residue that remains constant for most species, when and if you find one where it is different, that sequence also has other changes in it. Some of those will be compensatory. Originally neutral mutations which, once they had happened, allowed that ultra-conserved residue you thought was essential, to also change.
Sure. Apparently it’s confused some people. Other people have not been confused by it and are even using it with nary a word from you about it.
I’m guessing I was using it because Rumraket used it in his post on how to calculate FI.
What term would you prefer?
I would imagine hanging about at UD as you do you also get that feeling there too. I guess that makes you feel special, that you know what’s going on on both sides when so many are just cheerleaders, blindly following.
Sorry Rumraket, but we need to lose that “minimum threshold” concept.
🙂
Rumraket,
Talking about making shit up 🙂
I haven’t been following your argument I’m sorry to say.
It seems to me we can’t lose the minumum threshold, otherwise what is our fraction going to look like? What are we supposed to put as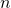 in
in 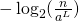 ?
?
I like the visual metaphor of the threshold represented by the plane dissecting the cone from Szostak 2003:
Rumraket,
I have compared multiple species Rum. Please get clarification before you ramble nonsense.
Rumraket,
Look at the 🙂 Rum. He was pointing to Jocks argument.
Have you not ready any of the papers I have linked or quoted you in this thread? Is this you just acting out some last ditch denial or what? You want a list of references?
How about you just start here:
Pervasive contingency and entrenchment in a billion years of Hsp90 evolution.
Or here
Alternative evolutionary histories in the sequence space of an ancient protein.
Or here
Epistasis as the primary factor in molecular evolution.
Or here
Pervasive cryptic epistasis in molecular evolution.
Or just go on google scholar and write ‘protein epistasis’ and read.
The choise is yours now. Continue to be ignorant and just brainlessly deny real world facts the way you do, or change your tune.
Rumraket,
Mung’s problem (well, one of Mung’s problems) is that he is treating the threshold value as an attribute of the ‘system’ as a whole. In Mung-world, a system has a single threshold value, and a single FI value is calculated, for the system as a whole. There cannot be different sequences that have different FI values.
If the threshold is set equal to the minimum activity level, then ALL sequences in the sequence space now have zero FI.
Yep, it’s that simple.
E4typo
OMagain,
And unnecessary when you have the best coverup artists on the planet scanning the comments 🙂
Rumraket,
Once you create a convincing argument that the FI in all proteins are the same or even similar. Without this you can cite papers until the cows come home but you won’t be supporting your argument.
Literally everything I wrote is true. I don’t care whether you did seven or ten pairwise alignments. Nobody cares.
It’s a meaningless exercise that doesn’t actually tell you about how conserved the protein is across the history of diversity of life. Or whether and/or to what extend epistasis is shaping the protein’s sequence or evolution. At minimum you need to do a multiple sequence alignment, and you have to sample much broader taxonomic diversity. And then you need to look at your results and think about what is going on, rather than just cite me what percentage of residues are identical between five to six cases like human vs e coli, fish vs bird, and pig vs ant.
Nobody is impressed by your ability to copy-paste long lines of text into a browswer window, press the button that reads “align”, and then ready out the “identity” field.
I know buddy. And I get that I was saying something that DNA_Jock seems to have contradicted. Unlike you I don’t mind if I end up disagreeing with someone on “my team” and I don’t need to go ask them for their input to save me from hard questions. Unlike you I can think for myself, and when I fuck up I can admit to it too.
I like to too. But someone might confuse the plane in the diagram with the line in the fraction and get all confused. They might think that a^L are below the plane and that n is not included in a^L and all other sorts of weird stuff.
They might even think that anything below it has no function at all!
Yep, it’s simply that false.
I have repeatedly stated that the threshold is arbitrary and can be set where anyone likes. Also, that as one relocates the threshold that the FI will change. I guess you missed that.
So your fictional Mung-world carries no resemblance to what I have actually written or to what I actually think. Please do better.
Who the hell said anything about FI in all proteins being the same or similar? What would that even mean? Nobody has made that claim and it isn’t necessary to make a claim like that to show that epistasis can potentially open up otherwise conserved residues to change.
I brought up epistasis in proteins. Epistasis. The phenomenon where multiple residues in a protein contribute to the same function, and where they can potentially compensate for each other. An amino acid position (in a protein sequence) that otherwise seems unable to change to another amino acid, ends up being able to if one or more other amino acids in the protein have changed first.
You responded to me talking about epistasis, and the claim I made that the phenomenon is ubiquitous in protein structure, function, and evolution, by saying that I was making shit up.
I then went and gave references (even though I already had previously in this thread, which your brain must as usual have somehow erased). And now you’re suddenly drooling about a claim nobody has made. But I see you have elected to just ignore the references. I guess that makes it easier to argue for the position you take. Who cares about pesky real-world experiments? Not you, that’s for sure.
Rumraket,
This assumes that there is no design modification between species. Other then assertion you have not explained why 5 or 6 species is not representative of how well preserved the sequences are.
Rumraket,
I don’t care when they don’t support your claim which is that I can do an experiment on protein A and understand the FI of protein B or protein complex C. What does the FI of one of Thorton’s enzymes have to do with the beta chain of ATP synthase?
I have made no assumptions about what constraints are operating on the protein in other species. Anywhere. I’m not claiming that any particular change we see between species is without some potential functional consequence. That’s just plain false Bill.
Uhh how about the actual demonstration with an alignment of about 100 species I did earlier, sampled from across the diversity of life?
Here’s a new one with a few more: https://www.uniprot.org/align/A20180705A7434721E10EE6586998A056CCD0537E833C950.
~12.1%
Where are these people claiming that the FI of protein A tells us about the FI of protein B? Where? Quote them.
Rumraket,
If you are not making this claim then I apologize for my misunderstanding.
Rumraket,
Thanks for the link. I played around with it and it appears a lot of the variation is around different types of bacteria. If you blast 3 different type of bacteria you can get as low as about 50% agreement. If I blast sharks, humans, mice and worms I get almost 80% sequence agreement. If I drop worms it is almost 90%.
No it isn’t. As I said, the “inference” only points to your misunderstanding and your willingness to forget any standards when it comes to your preferred “explanation,” while keeping an excessive skepticism when it comes to your least liked explanation. When it comes to nature you keep extreme skepticism to any evidence. When it comes to gods, anything does. Do you really not see the problem here?
No it doesn’t. It just works by using the very logic you claim for yourself, only used properly. You prefer to forget the requirements for design to be produced in order to conclude design. I prefer not to ignore the obvious. Design requires nature to have energy flow available. It requires beings made of the very things you claim to be designed. You cannot cherry-pick from designers as you please.
Nope. Inductive reasoning is pointing us to nature first, designers much much much later as a tiny tiny tiny product of an enormous nature. Think about it carefully Bill. You keep asking us to take all those examples of FI being produced by design, but you want us to dismiss the many requirements necessary for that design to be possible. Sorry, but that’s not inductive reasoning. That’s poor science and poor philosophy. And I’m being very kind with those adjectives.
I see our deep and undeniable dependence on nature. You don’t. Why not? because you want your conclusion to be true. No other reason.
Until you can explain that enormous gap in your standards towards either “side,” you cannot expect to gain much credence among the better informed. I’d expect though, that you should understand that our position is reasonable, while yours depends on having that enormously polarized double standard, and thus, it’s far from objective. It will never convince the philosophically and scientifically informed.
Entropy,
I think we have very different positions here. I don’t see any other reasonable explanation for FI. The rest of your philosophical discussion is interesting but for me the very powerful design inference trumps it.