Dr. Winston Ewert put forward his module hypothesis, but I put forward an alternate module hypothesis at the domain and motif level of proteins. It is based actually on papers by evolutionists who have pointed out that the problem of “Promiscuous Domains” remains an unsolved problem in evolutionary biology.
When I put Promiscuous Domains on the table in the Common Design vs. Common Descent thread, the TSZ Darwinists ignored the problem and then declared victory. I viewed their non-response as evidence they didn’t understand the problem and/or preferred to ignore it.
Perhaps pictures are worth ten thousand words. From the NIH, that great source inspiration for the Intelligent Design community, we have the CDART database viewer.
From the CDART viewer, I provide a few of the thousands of diagrams that show the promiscuity of protein domains. The diagrams below show the classical zinc finger ZF-C2H2 “ZF” domain and the Plextrin Homology “PH” domains. Note how the location of domains is “shuffled” to different locations in different proteins. It’s as if proteins are made by different lego-like parts in different order and position. My preliminary look into small 4-amino acid motifs that are the target of phosphorylating kinases suggests the the problem of promiscuity goes all the way down to small motif levels.
Such promiscuity is more consistent with common design than common descent.
Click to Enlarge Classical ZF-C2H2 Zinc Finger Page 5
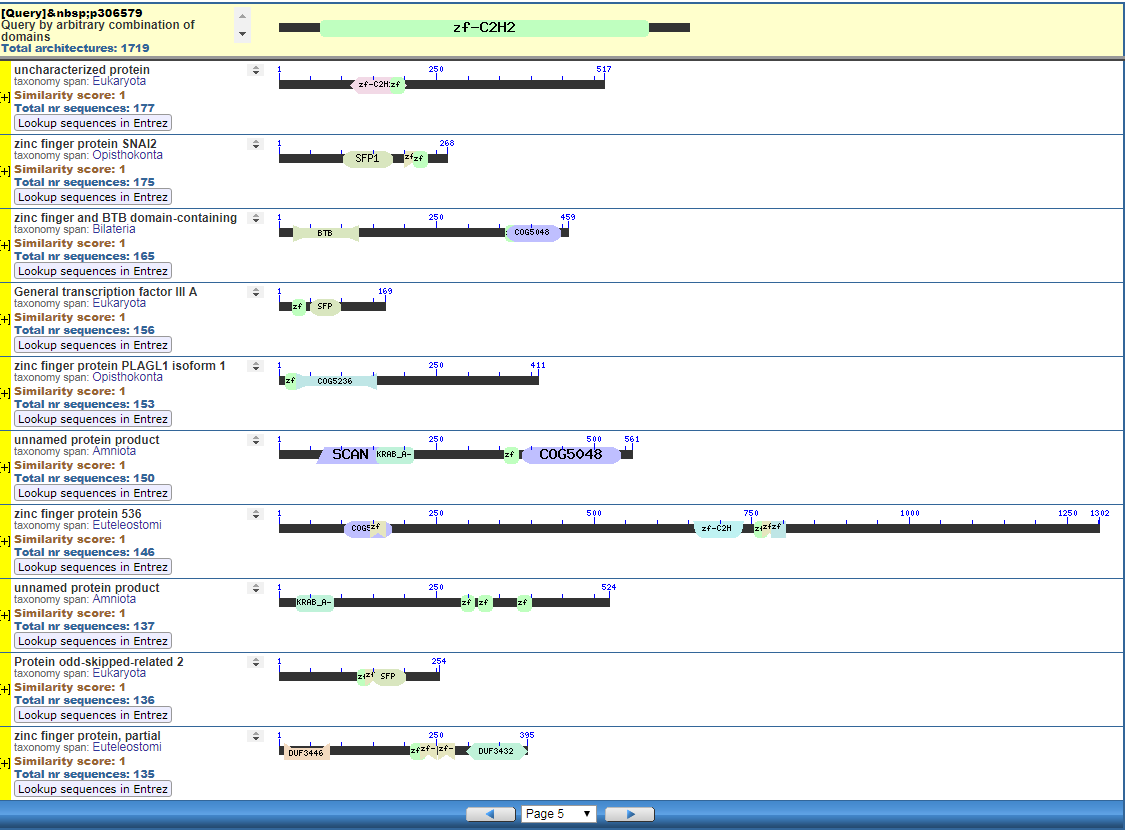
Click to Enlarge Classical ZF-C2H2 Zinc Finger Page 157
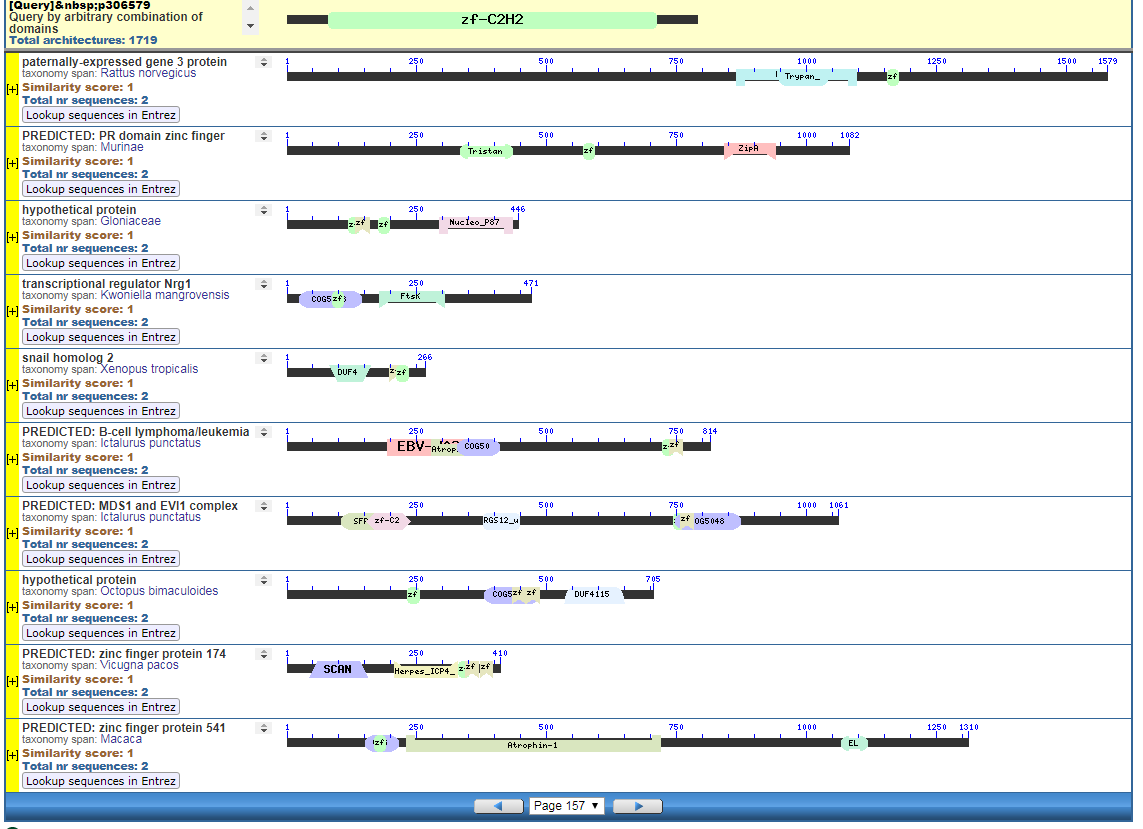
Click to see all CDART Classical ZF-C2H2 Zinc Finger Architectures

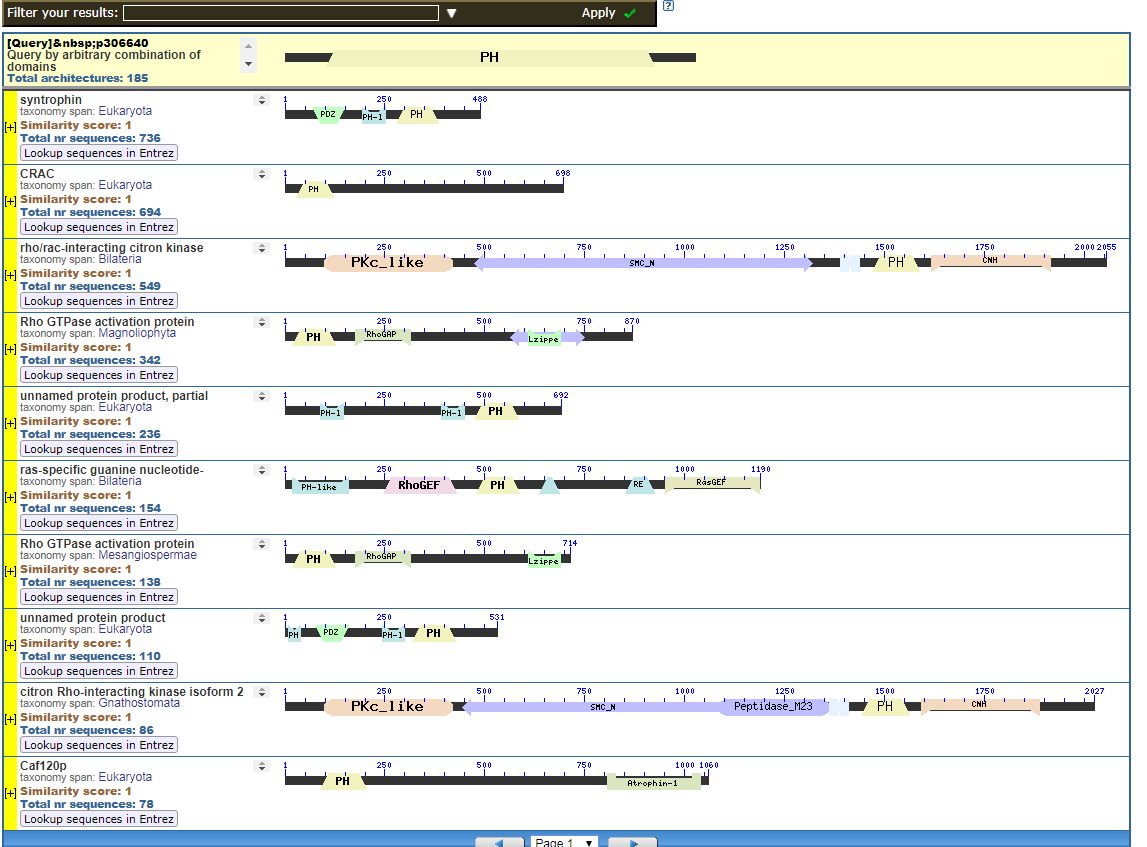
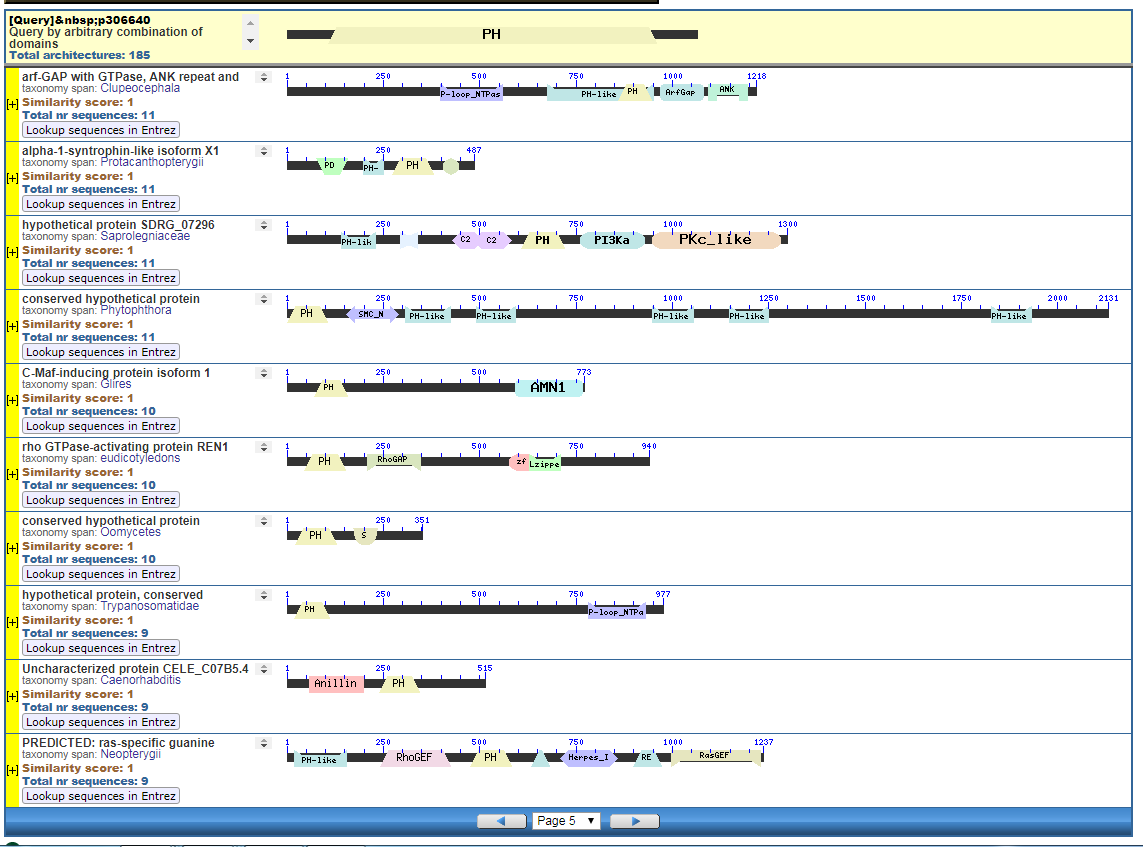
Mung,
Tomayto, tomahto.
I did.
If it is possible at all.
If knowledge is a justified true belief and and the only way to objective truth is divine revelation, then the way to knowledge is by the objective truth of the belief being justified by revelation. Knowledge about the world requires revelation. How this revelation works was my question.
Discovery is the gaining new knowledge. Without justification it is not knowledge. Without revelation it is not justified.
Seems like the same subject
How do you know which your favorite is?
If one can be mistaken about one’s knowledge about the existence of a deity, can one be justified in the knowledge of one’s favorite ice cream?
You can probably find the code for it online.
No need ,one must merely presuppose it.