Dr. Winston Ewert put forward his module hypothesis, but I put forward an alternate module hypothesis at the domain and motif level of proteins. It is based actually on papers by evolutionists who have pointed out that the problem of “Promiscuous Domains” remains an unsolved problem in evolutionary biology.
When I put Promiscuous Domains on the table in the Common Design vs. Common Descent thread, the TSZ Darwinists ignored the problem and then declared victory. I viewed their non-response as evidence they didn’t understand the problem and/or preferred to ignore it.
Perhaps pictures are worth ten thousand words. From the NIH, that great source inspiration for the Intelligent Design community, we have the CDART database viewer.
From the CDART viewer, I provide a few of the thousands of diagrams that show the promiscuity of protein domains. The diagrams below show the classical zinc finger ZF-C2H2 “ZF” domain and the Plextrin Homology “PH” domains. Note how the location of domains is “shuffled” to different locations in different proteins. It’s as if proteins are made by different lego-like parts in different order and position. My preliminary look into small 4-amino acid motifs that are the target of phosphorylating kinases suggests the the problem of promiscuity goes all the way down to small motif levels.
Such promiscuity is more consistent with common design than common descent.
Click to Enlarge Classical ZF-C2H2 Zinc Finger Page 5
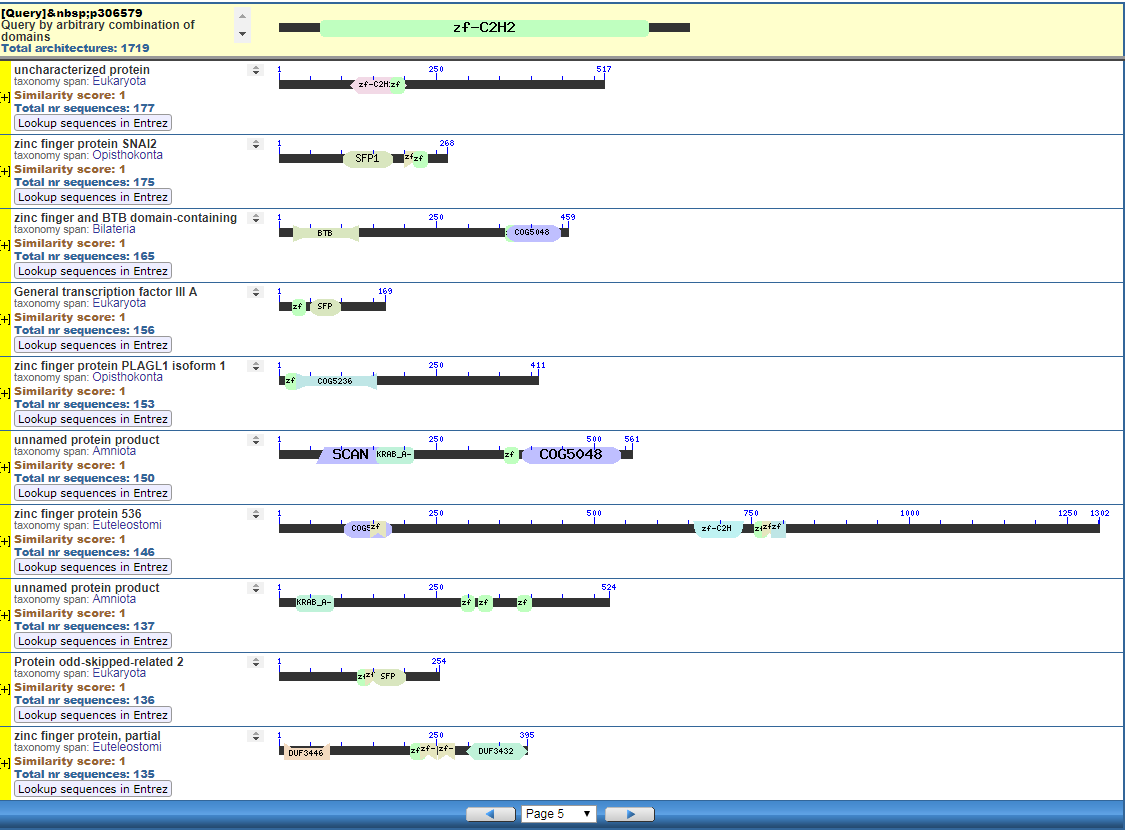
Click to Enlarge Classical ZF-C2H2 Zinc Finger Page 157
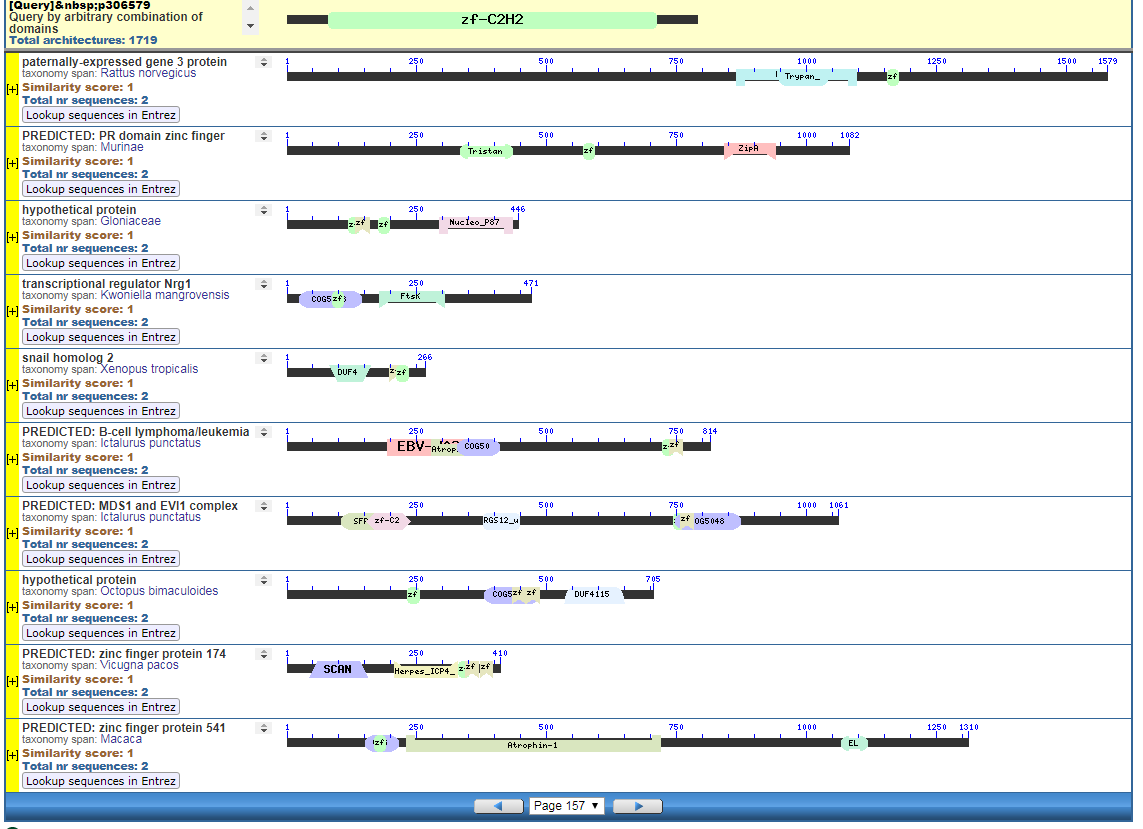
Click to see all CDART Classical ZF-C2H2 Zinc Finger Architectures

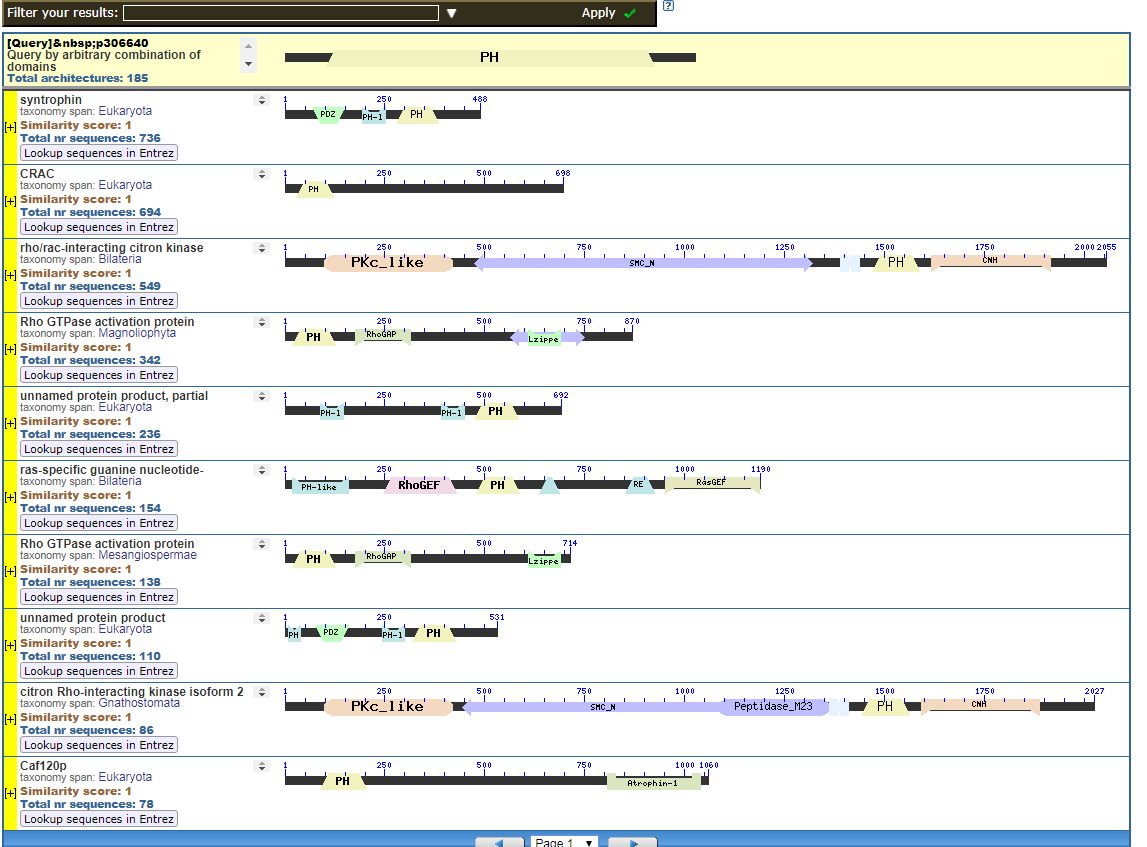
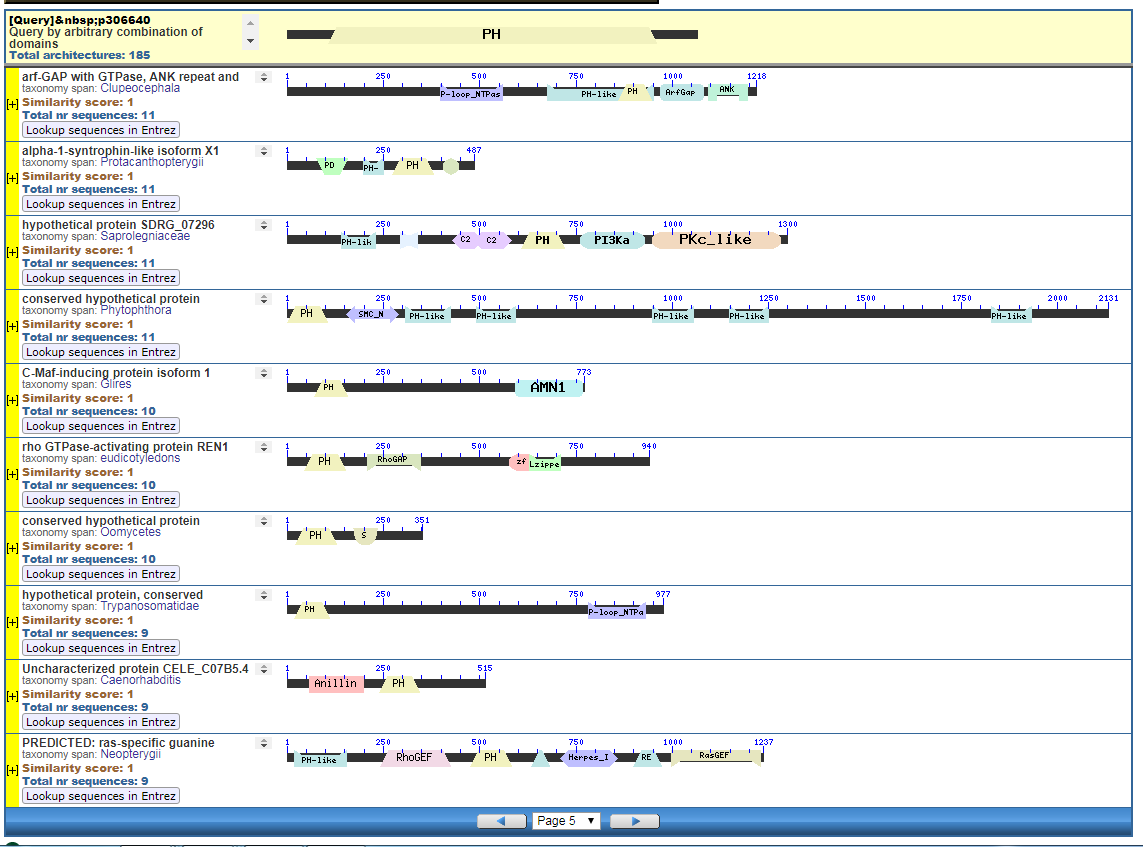
How you got to that conclusion from what I wrote, only the FSM knows.
I would want to have evidence to believe something exists, but the way God is defined, there can’t be any evidence of its existence, because EVERYTHING, by definition, exists because of God.
And I reject that anyway because it so obviously begs the question and it’s useless to explain anything at all
But again, wasn’t it you who was looking for the proverbial Jesus-face-in-a-bread-toast by looking at patterns in DNA and whatnot?
WTF? Have you been possessed by FMM or what?
Pain is a sensation, why should you know that absolutely?
This makes little sense. First of all it seems to me that nothing is certain, and we are not in a position where we can claim to know anything “absolutely”. But I also don’t think that having absolute certainty is a requirement to be able to live, function, think, and do science. We can get by with having tentative estimations of relative likelihoods. On the evidence, A seems more likely than B for example. So we go with A until new evidence potentially tips the scales in favor of B.
That is acting on the situation before us and the information we have. That isn’t certainty, far from it, but that just (at least in my view) goes to show that certainty is not a precondition for life, or acting rationally.
I am relieved to hear it.
You certainly can’t build a phylogeny of proteins, even if all proteins really did commonly descend from one. An example I already mentioned: the Class l and Class ll aaRSs bear (almost) no relation to each other. If they share common ancestry, that information has been obliterated.
Isn’t it true that, even if protein evolution could be traced back to a single, universal common ancestor protein, that organism would predate LUCA by quite a lot, and therefore Sal’s claim that UCD is in question unless we can figure out how proteins evolved before LUCA is nonsense?
I mean, even if LUCA was poofed into existence with a library of proteins from which all the extant ones evolved, that would still be UCD
OK, I guess that’s wrong (because of those taxonomically restricted proteins) but you get the point
Yes, if the proteins found in LUCA themselves share a genealogical relationship from a common ancestral protein, that would imply LUCA was not the “first organism” but itself had a long evolutionary history of ancestor species in which this repertoire of proteins evolved. That is indeed what we find by the way. Multiple proteins and RNAs found in the translation system are themselves genealogically related, and therefore indicate that there was a substantial period of pre-LUCA evolution. LUCA was not the first organism to appear immediately following the origin of life. There was a substantial period of evolution before LUCA. Perhaps as much as 800 million years.
Yes. In that case all proteins could be traced to the same common ancestral species, not the same protein.
Yep, LUCA is effectively the coalescent of a collection of gene trees, whereas each individual gene must logically coalesce further back (the coalescent being, to my biochemically addled mind, the DNA template from which any given set ultimately descends.)
Rumraket,
Allan Miller,
Thanks guys
Sal,
This reads as if you have some very deep conceptual problems. It doesn’t make much sense.
1. What do you mean by “learning pFam”? The wording seems rather weird when talking about a database of domain models.
2. Each square in that schema represents a domain. Some of those domains are homologous to each other (they’re thus drawn with the same color), which means that they share common ancestry. Therefore, it doesn’t make sense to look at this and say “there might be a unifying architecture.” What do you even mean by that? How could looking at proteins containing KRAB and Zinc finger domains mean that there’s a unifying architecture? Do you think that every protein has these two domains? If not, then what do you mean by unifying architecture?
I’d ask you to consider that you might have very deep problems in your understanding of many, but many, things, but we both know you’re unlikely to read this far, and pretty much unable to consider that you might have misunderstood something.
One more that will go unread.
Nope. I’m not saying such a thing. I know of nobody who’s saying such a thing.
While I never advertise them, I have built protein phylogenies. Before doing so, I have gathered proteins that belong to a single family, meaning they’re homologous to each other. That alone means that I don’t try and build phylogenies for proteins that are not clearly related, and thus that in building phylogenetic trees for a subset of a protein family I don’t assume that all extant proteins share common ancestry. Therefore you’re going for a non-sequitur.
What? besides this being a non-sequitur, since it would be surprising if scientists were making phylogenetic trees for proteins with different domains, all that’s needed to make new protein architectures is recombination. There’s nothing magical about it. I suspect now that you don’t know what “protein architecture” means.
So what?
Sal,
I’m not trying to be insulting here, but you’re really convincing me that you are really off your element here. But authentically far off from it. Your comments don’t make much sense. They show deep problems in your misunderstanding and astounding incompetence. You’re therefore ridiculing yourself.
Just so you know.
I think you misunderstand what “universal common ancestor” means, possibly on purpose. You actually admit ahead of time that nobody is proposing that a single protein is the universal common ancestor of all proteins, so failure to find such a thing is pointless.
Further, you ignore the fact that there are ways to evolve new proteins that don’t require them to be descended from previous proteins. Are you aware of any of these ways?
People don’t want to discuss things with you because you treat TSZ as your personal brain dump.
What two architectures? Where do you show these two architectures? All I see is pretty pictures with illegible text.
And that’s all the audience in the church basement will see as Sal gives his presentation. However for them the idea that those pretty pictures and illegible text represent actual science will be true, not false.
Freaking studly good insight! Thanks.
Nevermind, Allan Miller gave me some good data on the architecture of aaRS proteins. Class 1 vs. Class 2. Besides the consensus here so far is :
1. major protein families had indpendent origins, or at the least not some smooth traceable phylogeny like what we can do within a single protein family (i.e. Rhodopsins are within the family of Opsins).
2. even if hypothetically there were such a single protein from which all others descended, the phylogenetic signal has been obliterated
Ah, now you’re seeing the light. We don’t need to assume all proteins descended from a common ancestral protein to be able to do science do we. They also implement a nested hierarchy, by the way. But if one doesn’t need to assume universal common ancestry for proteins, why do we need the assumption of universal common ancestry for anything else to do science. The only thing we need to assert is “similar things may be have similarly.”
I’m mean, did all Zinc Finger domains descend from a common ancestor, or did they converge if they are found in different protein families? Was it gene fusion, exon shuffling, who knows? Who cares? Only evolutionary biologists, who’ll only say they don’t know anyway!
OK then. Don’t assume anything, you don’t have to to do science. Just let the data show you the truth
That’s your problem right there. IDists don’t care about explaining stuff, only real scientists do
IDists care about real science, Phylogenies that can’t be proven, where the holes in critical theories are filled with lots of “I don’t know” — this is not quality science.
Quality science are things like Quantum Mechanics and Electro Magnetism. That’s the real deal, baby.
NOTE: I’ve already said I don’t defend ID as being science. If push came to shove, I’d say ID isn’t science. But just because ID isn’t science, doesn’t mean IDists don’t care about science.
What a strange question. First of all common ancestry is not assumed, and nobody claims it has to be true for us to be able to do science. It’s just a conclusion drawn from looking at the data. We want to know what the facts are and we have an interest in our own origins and that is good enough.
Ahhahaha that’s funny coming from a guy who wants to replace science with “magic man did it” and zero way to experimentally demonstrate that. We always get this double standard from IDcreationists. You like to blather endlessly about wanting to see things experimentally demonstrated in front of your own eyes, otherwise it’s “phylogenies that can’t be proven”, but in their stead you would substitute in a blanket statement that an immortal, immaterial, omnipresent, middle-eastern bronze-age carpenter magically wished life into existence. But how do you even test that? How do you prove that?
Don’t give us this nonsense. You couldn’t give any less of a shit about what is good or bad science, you just want to evangelize and advertise for your religious beliefs.
But your solution is to replace the “I don’t know” with “MAGIC MAN DUNIT”. It’s not even bad science. It isn’t AT ALL science.
Take your cheap gig somewhere else, you’ve been had.
They did not abandon the old paradigm they simply noted the slight anomaly and went about their business.
Newtonian physics still works well for most things. It was not until Einstein came along that folks realized that their assumptions were irredeemable at certain resolutions .
That is exactly what I’m talking about.
When data seems to conflict with what we already “know” we don’t abandon our models and start from scratch. We try everything else we can think of first and as a last resort tweak our models as minimally as possible to account for the new observation.
None of this is meant to be a criticism. If we abandoned our assumptions every time things did not work exactly as expected science would never get off the ground.
I think it’s nothing short of a miracle that the universe is such that we can go at it like that.
It is certainly logically possible that the universe could have been an all or nothing proposition in which we either
know everything whatsoever or we know nothing at all.
peace
That claim is simply not warranted.
Actually, they continued to try and resolve the anomaly. “They did NOT say, ‘oh that’s noise, let’s ignore it.’” In fact, the attempt to resolve such anomalies is often the spur to new science, whether it is the discovery of a new planet, or a new theory.
Now, backing up a bit:
Observations which can be independently verified are construed as scientifically objective. Whether you like this sense of the term, it would behoove you to understand that that is how many people use the term.
We can determine the fit of data to a model objectively. For instance, Newtonian Mechanics is a better fit to the data than Aristotelian mechanics. In this case, objectivity can be defined mathematically.
You can certainly treat gray fur as a trait, but when looking at all observable traits, you will find that heritable traits form a nested hierarchy, in particular, gray foxes and red foxes have far more traits in common than gray foxes and gray squirrels.
If you want to use the term in that idiosyncratic that is up to you but then the entire argument that the “objective” nested hierarchy is strong evidence of common decent simply evaporates.
Not unless you twist the term to mean something other than objective. If you only mean something like “repeatedly” or “popularly”.
Then yes you can determine that the data fit in that way.
Any categorization will form a nested hierarchy. That is how categorization.
That is simply not true. The only way that works is if you weigh certain characters more strongly than others.
Characters are not like items on a Chinese menu they don’t form a list that you check and then tally up the total they are inherently subjective.
If you provide a list of things that you feel are more similar in grey and red foxes I can provide a longer list for grey and red squirrels.
If my list is shorter than yours at first I will just need to look harder.
peace
What would such a universe look like?
I think Zach would agree, grey and red squirrels have more in common than grey squirrels and grey foxes.
The concept of “scientific objectivity” is hardly idiosyncratic. While perfect objectivity is not attainable, the scientific method is designed to minimize individual biases.
That is incorrect. Branching descent with modification predicts a nested hierarchy, and that is the pattern we observe.
In the case of Newtonian vs Aristotelian mechanics, we can apply mathematical principles comparing the theories to the data. Newtonian mechanics more closely matches observations.
Not all categorizations are hierarchical, but anything can be put into a nested hierarchy. However, if we group organisms by observable and heritable traits, then they fall into a nested hierarchy pattern.
Weighting isn’t required. Just put all the traits on the table. Gray foxes and red foxes have more traits in common than squirrels of any color. For instance, you could look at the DNA sequences, which can be mathematically analyzed, and that would show more similarities between gray and red foxes than squirrels of any color.
Well, gee whiz. You’re right! Gray and red squirrels have more traits in common than foxes of any color. Consequently, we group squirrels together and foxes together when considering all observable traits.
Of course not! When confronted with things they don’t know, scientists should say god-did-it instead. That would be hyper-high-quality science indeed.
Of course! In those fields no scientists has ever said “I don’t know”! They always say “god-did-it” when confronted with things they cannot explain.
like a computer program that requires a password to access or a combination lock on a treasure chest or a secret message that looks scrambled until you apply the key to unlock it.
There are lots of things we encounter in life that are information rich but look like gibberish to the uninitiated.
peace
How exactly do you propose we do that??
Do you actually think there is an approved master list of traits out there somewhere.
The more you observe the more traits you will discover. If we look closely.
If you choose you can go down to the quantum level or out to include the context of entire environment or forward and backward in time to include all of the history of the organism to find distinguishing qualities or characteristic (traits).
LOL nothing is more funny than laughing at folks unintentional fopaux instead of reading and interacting charitably.
It always good for a laugh 😉
peace
Sure or you could could look at something like the propensity and ability to climb trees which grey squirrels share with grey foxes.
We can mathematically analyze that as well
grey foxes have a 100% similarity with grey squirrels in this regard and a 0% similarity with red foxes. This is a much more pronounced result than you get with comparing DNA mathematically.
Or you could choose to look at preferred habitat. Grey squirrels and Grey foxes tend to inhabit denser forests while red foxes tend to inhabit more open habitats.
If we wanted we could do this mathematically as well I suppose we could rank the likelihood of finding a grey fox in proximity to the open habitat red foxes frequent.
I could go on and on, but if you don’t get the point by now it’s unlikely you ever will
peace
Also Gray Langurs, gray climbing frogs,gray climbing mice, wolf spiders can be gray, humans can climb trees and have gray hair, cats are also know to be gray ,Care to construct a nested hierarchy with those members?
Sure, place all the gray tree climbing things with regard to the habitat and put then in the hierarchy and see if the trees match.
Actually gray foxes have quite a bit of red fur, so if color is relevant red foxes and gray foxes have quite a bit in common. In fact gray squirrels can have brownish fur.
Is the mechanism of that squirrels and foxes use to climb tree the same? Same as the Langur, tree mouse ,cats?
Depends on whether you’re talking about Western gray squirrels or Eastern gray squirrels (two separate species, though FMM may not think so).
Such as, for example, the field of molecular phylogenetics.
Make a list of observations. Or, in the case of DNA, the DNA sequence is the list.
That’s right.
If the behavior is heritable, then sure, you can list it. Now what about the hundreds of other phenotypic traits (e.g. teeth, ankles) and the thousands of genotypic traits? When we look at all the traits, which grouping is better?
{{red foxes, red squirrels} {gray foxes, gray squirrels}} or
{{red foxes, gray foxes}, {red squirrels, gray squirrels}}
Objective truth rears its ugly head yet again.
He does seem to have missed that he did have expectations and that what he witnessed went contrary to those expectations.
That’s not what he said.
Yes. Gene duplication followed by divergence.
The alternative is that novel proteins appear magically from the primordial DNA soup. And you may as well be a creationist if you believe that.
Mung,
Was this before or after the origin of the ribosome 🙂
Are you familiar with the concept ‘false dichotomy’?
Allan Miller,
Is that the opposite of a true dichotomy 🙂
No ,actually many things are funnier than treating a faux pas as intententional. You just never know because people charitably never mention it.
FMM’s knee-jerk misapprehension that I am claiming to be bias-free is leading FMM and Mung astray. I made no such claim. My point was that the consilience arises from the data, and NOT the group-think. Scientists who prove the group-think wrong get Nobels. But FMM was claiming that any data that do not fit the group-think are discarded as “noise”. He’s wrong about that.
Hardly.
I wrote:
Do at least try to keep up, Mung.
Don’t think so, each time you tried a password and it failed ,you would know it was not the password.
I have been thinking, in a Universe where there is only one thing to know might fit the requirement.
I agree. Then again there are things that are just gibberish.
peace
Really, fifth believes humans can know and discover things sans revelation?
Ask fifth.
What he wrote was:
You changed the subject to “discovery and knowledge.” So no, I don’t think he is saying you cannot discover what your favorite flavor of ice cream is or know what your favorite flavor of ice cream is.