Background
For the past month or so I’ve been investigating the claim that the phylogenetic signal is evidence that a dataset shares common descent.
Supposedly, the phylogenetic signal is one of, if not the, strongest pieces of evidence for common descent. It is one of the first of the 29+ evidences for evolution offered over at Talk Origins (TO).
Quoting from the article:
The degree to which a given phylogeny displays a unique, well-supported, objective nested hierarchy can be rigorously quantified. Several different statistical tests have been developed for determining whether a phylogeny has a subjective or objective nested hierarchy, or whether a given nested hierarchy could have been generated by a chance process instead of a genealogical process (Swofford 1996, p. 504). These tests measure the degree of “cladistic hierarchical structure” (also known as the “phylogenetic signal”) in a phylogeny, and phylogenies based upon true genealogical processes give high values of hierarchical structure, whereas subjective phylogenies that have only apparent hierarchical structure (like a phylogeny of cars, for example) give low values (Archie 1989; Faith and Cranston 1991; Farris 1989; Felsenstein 1985; Hillis 1991; Hillis and Huelsenbeck 1992; Huelsenbeck et al. 2001; Klassen et al. 1991).
http://www.talkorigins.org/faqs/comdesc/section1.html#nested_hierarchy
I’ve been skeptical of this claim. A tree is just one kind of directed acyclic graph (DAG), and my hunch is many kinds of DAGs will also score highly on metrics for phylogenetic signal. I picked one metric, the consistency index (CI), which according to Klassen 1991 is the most widely used metric. It also is the featured metric in the above TO article. Plus, it is very simple to calculate. So, I’ve focused my efforts on the CI metric.
Result
What I have found is that my hunch is correct. It is simple to create a DAG that scores highly in CI, well within the range of published CI scores for real datasets.
Consequently, it is incorrect to say the phylogenetic signal is strong evidence for evolution. In particular, this claim is provably false (as I have proven here):
Phylogenies based upon true genealogical processes give high values of hierarchical structure, whereas subjective phylogenies that have only apparent hierarchical structure give low values.
http://www.talkorigins.org/faqs/comdesc/section1.html#nested_hierarchy
How have I proven it false? I generate DNA sequences from directed acyclic graphs, and the trees derived from these sequences using well established methods produce CI scores well within published ranges. Here are two such experiments plotted on the chart from Klassen 1991:
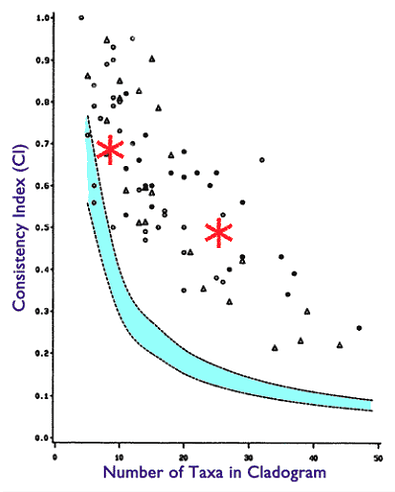
This is a phylogeny with very high value of hierarchical structure not generated from a true genealogical process.
Methods
To reproduce my results you can run the DAG dataset generator here: https://repl.it/@EricHolloway/Phylogenetic-Signal-Fallacy-Nucleotide-Level
Take the generated DNA sequences, which are in FASTA format, and paste them into the ClustalW online tool.
Take the results of the ClustalW tool, and use the PAUP software to generate trees and measure CI scores. You’ll need to fiddle with the NEX file format, so to save you the trouble, I’ve included an already created NEX file that I’ve generated from the aforementioned process, which you can pop into PAUP.
Once you load a NEX file into PAUP, here are the steps to generate trees, and then measure CI.
- Press “Generate Trees” in the “Trees” menu.
- Press the “OK” button.
- Press “Describe Trees” in the “Trees” menu.
- Press the “Describe” button.
- You will see something like the following:
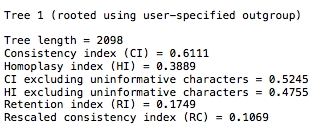
You will find the 27 taxa in the file will generate CI scores in the range of 0.48-0.53. If you look at the Klassen 1991 chart, you will see this is well within published scores for that number of taxa.
Conclusion
So, what is my takeaway from this?
Basically, highly statistically significant CI scores do not indicate common descent. They can just as easily be generated by a DAG. Therefore, we cannot infer common descent from high CI.
Furthermore, insofar as CI is representative of the state of phylogenetic signal measurement, my result undermines the more general claim that phylogenetic signal indicates common descent.
As such, the Talk Origin’s claim that the nested hierarchy of species is well attested by the data is highly questionable if not outright false, and should be retracted as evidence for evolution until such time as a much more rigorous analysis with DAG eliminating controls is established.
Addendum
To visually illustrate what I mean by a DAG generating the DNA sequences, here is a graph of one such DAG. Each colored/numbered box represents a gene, which is replaced by a unique, randomly generated (uniform over ‘GATC’) DNA sequence of 20-30 letters long in post processing. Arrows indicate when ‘ancestor’ gene sets are combined into larger gene sets. If you look closely, you will see each gene set contains the union of all incoming gene sets, plus one new gene. As you can see, this looks nothing at all like an evolutionary process, yet it produces very high phylogenetic signal as measured by the consistency index (CI) metric.
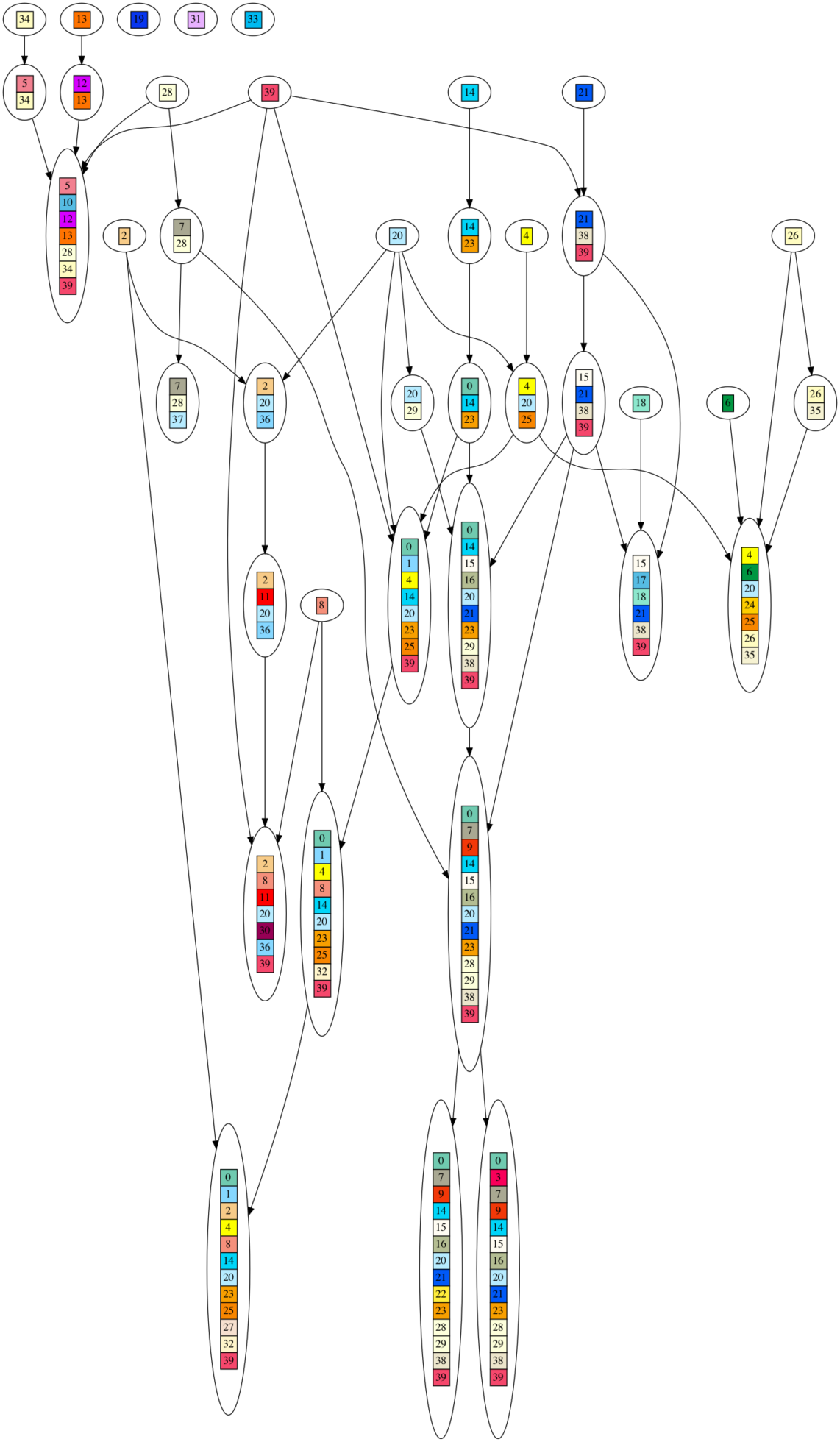

John Harshman,
Until there is a clearly articulated hypothesis there is nothing to test.
Experience shows that we can go around in these circles forever, with you never learning anything but continuing to post your one-line obfuscations.
John Harshman,
I honestly do not either know where this goes either. I think Behe’s design criteria is telling us where the edge of scientific inquiry is. We are no where on how to model biological innovation. We are also know where on how to model the origin of matter or atoms. Except for the time that YEC assigns to Genesis 1 it appears to be a pretty good guide on where science starts.
colewd,
“Any two species in the same genus are related by descent” seems a clear enough hypothesis to me. If we would accept that hypothesis as supported by a particular test when replacing ‘genus’ by a lower taxonomic level, we’d be a bit inconsistent in rejecting it at the level of genus, for some arbitrary reason. Reductio ad absurdum, the consistent position, then, is to reject all such tests, including forensic analysis, food composition and contamination analysis, and genealogical tests. It’s remarkable what can get thrown out with the bathwater when you really dig your heels in.
What do you mean “either”? I totally know where this goes. You have been following the same script for years, and it leads nowhere.
I speculate that you have put your finger on one of the key strengths of the magical sky daddy poof “explanation” – it explains absolutely everything, is consistent with any possible observation, and cannot be demonstrated to be wrong. The hypothesis that X can ONLY result from process Y is useless by definition, since poofing can produce any X.
Which comes in handy when we don’t know how X might have resulted from processes not understood. Poofing always suffices. For some folks, it’s perverse to look for some other process, since we already have one fully sufficient, and another proposed process can only create confusion and is probably not foolproof.
Flint,
There would remain a nagging question-mark over my cartoon head as to why, to generations of biologists, it all looks like descent, in a very detailed manner. It’s not the first time I’ve said this, but if someone has gone to such extraordinary lengths to cover their tracks, maybe they don’t want to be seen? The paradox is that believers, having invested remarkably little effort in grasping the evidence despite many years of engagement in some cases, can’t even see the subterfuge!
I’ve never seen evidence that you’ve looked at anything very closely — not even the ID literature. You flit from one topic to another, apparently believing that your combination of a biblical worldview with superior powers of ratiocination enables you to see immediately what people who’ve worked for years to acquire specialist knowledge have gotten wrong (presumably because they are committed to a materialist worldview).
A legitimate scholar would first do an extensive review of published analyses. If you were to do that, then you might learn how to do a correct analysis yourself. Instead, you proceed under the false assumption that you’re equipped already to make novel contributions to whatever field captures your fancy. That’s magical thinking, Eric — magical thinking about your own capabilities. It’s truly a sickness.
Is that an electroencephalogram on your head, Eric?
Allan Miller,
How would you test the hypothesis that two species of the same genus share a common ancestor? At what point does the test fail?
Don’t let them discourage you, Eric. Of course there is no “evolution”. They can’t even reply with their own “fitness” function, so their very foundation is a fantasy.
What about abiogenesis? Oh, wait.
Consilient, eh? And swathes. Wow!
Yeah, Bill. Accept first that “evolution” is true, and only then test if “evolution” is true? How could you miss that?
Yep. Science is all about voting. Except when ALL knew bloodletting was the best. That was bad. But biologists are not much into science, so they’re cool.
Yes, Eric. And write some damn equations even if they don’t mean anything. They’re cool and sexy. Chicks fall for them all the time. If you’re into chicks that is…
Enough with the silly questions. That’s your assumption. You still don’t get it?!?
How would you test the hypothesis that two species of the same genus do not share a common ancestor? At what point does the test fail?
You can at least test the hypothesis that a genus is monophyletic. If it is, that’s good evidence that they share a common ancestor. If it isn’t, that’s good evidence that if there’s a common ancestor, it isn’t the one you thought it was.
See, for example, Chesser R.T., Barker F.K., Brumfield R.T. Fourfold polyphyly of the genus formerly known as Upucerthia, with notes on the systematics and evolution of the avian subfamily Furnariinae. Molecular Phylogenetics and Evolution 2007; 44:1320-1332.
As I said in my original comment, I invite you to replace ‘genus’ by a lower taxonomic rank, and then answer that question for that rank yourself. If you reject phylogenetic inference at any level, for consistency you should reject it at all, right down to genealogy or disease tracing. If you don’t, where do you insert Colewd’s Razor, and why does the method stop working on the other side of it?
Eric already writes equations that don’t mean anything. That is precisely the topic of my post “Stark Incompetence.”
Allan Miller,
I am not rejecting phylogenetic inference. I am suggesting that the next step is testing a more defined hypothesis if that is indeed practical. It very well could be that this is not practical for very good reasons as if it was we would not be struggling so hard with this discussion.
colewd,
The only one ‘struggling with the discussion’ would appear to be you. I am happy to accept a high percentage of alignable sequence (say, above 50%) as indicative of common descent (strictly, common sequence descent) thoughout the ToL. You reject that, at some taxonomic level or another which you never articulate.
The only person struggling here Bill is you. Your willful scientific ignorance drags you down in every discussion you attempt. Or “discussion” since you never actually pay attention to any scientific evidence presented.
What does that mean?
Allan Miller,
Align-able sequences are an indication of common design also. How do you isolate the signal from the noise? Where did design stop and common descent take over? The YEC guys are trying to also figure this out as they don’t have clear definition of kinds at this point. Nathaniel Jeanson has provided some interesting data on this subject around mitochondrial molecular clocks.
Then I should pay attention the the meaningless equation war. Darwin’s prophecy of a Disturbance in the “evolutionary” Force followed by a total equation war is coming true. Is there anything that guy didn’t know?
colewd,
Among the morsels of word salad it’s possible to pull out a few things to respond to.
Why? Wouldn’t common design produce, if anything, identical sequences?
That’s some of the word salad that can’t be deciphered enough to respond to.
Have you ever wondered why it’s so hard to locate kinds? I could suggest an explanation if you like.
Has he? Are you sure? As far as I know he’s never provided any data, much less on the subject of kinds.
Doesn’t that clearly establish his bona fides?
Nonsense.
More nonsense.
Yet more nonsense.
Thank God that there’s no “fact checking” in place here at TSZ!
But 90% of the human genome is junk. Proof the designer is experimenting but likes to document failures. That’s why evolutionists had to invent “drift”.
Can’t check facts when you don’t include them in your “nonsense” assertions.
John Harshman,
In some cases we find identical amino acid sequences at the the class taxa level.
I don’t think it will be hard to get some level of stratification once the alignment data and gene data matures. Sal’s flower produces some level of stratification.
He has done interesting analysis from papers that have produced pedigree mutation data.
LOL! Sure thing Bill. We’ll have that genetic data for Creationist “kinds” any day now. Any day. 😀 😀 😀
You mean he cherry-picked a tiny bit of real data as support for his YEC woo. But we know telling the truth about science is exceptionally difficult for you.
Mung,
No, it’s true, I really am. It’s quite generous, really.
I disagree.
You tell me. I think ‘common design’, as an explanation for high sequence commonality, is bullshit; a feeble attempt to create equivalence – particularly when applied to sequence which (as far as we can tell) makes no difference in anything functional. There is no apparent need for squid succinate dehydrogenase to differ from octopus, for example. All it needs to do is dehydrogenate succinate. (And we should note here that you think functional sequences vanishingly rare in sequence space; how extraordinary that there are all these neighbours exquisitely varied according to organismal need!).
If you play the ‘you never know …’ card at this point, that would be special pleading.
Strange; it’s evolutionary theory that should have this difficulty, not discrete-instance no-change Creation.
Allan Miller,
Are you claiming cell division and sexual reproduction evolved?
Are you claiming that common descent is a good explanation for vastly different sequences providing the same function?
Yet the evidence sharply contradicts common descent as the cause of distinct family and classes of organisms.
http://www.sci-news.com/genetics/article01036.html
http://www.sci-news.com/genetics/article01036.html
Come back with those goalposts!
No, I’m claiming it is a good explanation for highly similar, but nonidentical, sequences.
Does it hell. How can sharing 70% of genes with zebrafish be evidence against a genetic relationship with zebrafish? Commonality is exactly what one would expect.
What would be interesting, though, is whether every gene, irrespective of function, clusters better with closer relatives than distant. I already know they will, without even looking. Ask me how.
Your little one-liners don’t actually say anything, Bill. At least try to make sense, if you can. Now, what proteins are you thinking of here? What “class taxa”? And most of all, how does that make any sort of argument?
Now that one was just word salad. I don’t think even a much longer explanation would make any sense out of it. But feel free to try.
But that isn’t data he presented, is it? And haven’t we already gone over his misuse of those pedigree rates? How is that even relevant?
John Harshman,
Alpha actin aligned in mammals both skeletal and heart muscle.
The gene sets identified in Sal’s flower indicates that zebra fish chickens mice and humans belong to different kinds.
I don’t agree with your argument. You label it misused but you simply disagree with his approach. Your disagreement does not equal misuse.
Stop with the one-liners. At the very least, cite something. What do you mean “aligned”? Are there different alpha actins in the two muscle types? What mammals are you talking about?
Why?
I didn’t make an argument.
John Harshman,
Heart and skeletal muscle have different AA sequences and have 100% alignment between chimps humans and mice. This is strong evidence the sequences were designed.
Different gene sets are evidence of unique designs.
Please cite something that shows this. Why is that evidence of design?
Please, no more one-liners. Explain and justify your claim.
Indeed. Then your designer, as you are, is a cunt.
A specific mutation in the AKT1 gene causes Proteus syndrome. That ‘gene set’ is designed, according to you. Therefore the designer designed that version of AKT1.
https://www.nhs.uk/news/2011/07july/pages/genetic-link-proteus-syndrome.aspx
It is believed that ‘The Elephant man’ suffered from Proteus syndrome. A unique design indeed. A cunt worshipping a cunt. How apt.
So if things are the same, that means design. But if they are different, that means design. So, finally, we have a good test for design.
Allan, most YECs understand now that Noah’s Ark, though incredibly large for a wooden boat, was rather small for a zoo. They have to posit a fabulous rate of diversification of types within created kinds of terrestrial organism, over the past 4300 or so years, to account for the present-day variety in types. As Mung once put it, YECs are evolutionists on steroids.
True, but kinds themselves should be discontinuous and hence easily defined, since each was a separate entity with no genetic connection to another.
In my opinion it is more sensible to think of Noah’s Ark as signifying the human being as a compendium of the animal kingdom as formulated in their own specific way by Lorenz Oken and others. It springs from the same imaginary thinking that produced the zodiac or animal circle that pictured the cosmic influences of separate animal types making up the complete human with each type relating to a specific body area.
The original story of the Ark was never meant to be taken literally. That is the modern way of thinking about things.
What about the Svalbard Vault?
Add to that the “types” and “variety” confusion, and suddenly meaningless equations remain you best bet.
Let me explain: look around and all you see is basic archetypes with lots of built in variety and adaptability (which btw is NOT “evolution”). So you don’t need to carry all the variants out there. Just have the one wolf pair and never worry about the dog or coyote, etc.
They ARE discontinuous. Here and now. And that’s really puzzling and CONTRARY to “evolution” given no “gradualism” and most certainly no “divergence of character” as discussed.
The genetic connection makes perfect sense on the account of common design. So no, genetics doesn’t support “evolution” ONE BIT more than Intelligent Design. In fact, genetic conservation is CONTRARY again to “evolution” which sure would have “evolved” all kind of new solutions all over if true.
Did you talk to the writer? How else would you know?
I am not trying to make novel contributions. I am merely trying to verify the fundamental premises of evolutionary theory.
My point here is I know at least one rigorous field decently, which I use everyday at work with a fair level of success and advancement, and have experience in a couple other rigorous fields, and in all such fields the fundamental results can be demonstrated conclusively to a layman without them first having to get a PhD in the subject. Give me anyone with basics of math and logic, and I can explain to them any result in Comp Sci that I know well enough. Even you have backed me, very reluctantly, in what I’ve said about halting oracles.
This is not what I experience in the slightest in my investigations into evolution. There is a lot of grandstanding, insulting, and otherwise demeaning and unprofessional behavior from the experts. And when I try to dig into the details of these claims, it all evaporates. It’s like there’s a Heisenberg principle for evolutionary theory, where a fundamental claim cannot be both clear and rigorously demonstrable at the same time.
So, whatever the level of veracity that lies in evolutionary theory, it is incorrect to say it is a scientific theory of the same caliber as fundamental theories in physics and computer science.
Do results from computional sciences often clash with certain religious beliefs?
Could you give an example of one, please? Should be easy, right?
You might consider taking a few courses in evolutionary biology first so you at least understand what it is instead of always attacking your ID-Creationist strawman. That however would take intellectual honesty which you have yet to demonstrate you possess.
“Investigation” 😀 😀 😀 To you that means reading the bullshit produced by the DI.
As always the combination of your brutal scientific ignorance and unwarranted arrogance never ceases to amuse. 🙂
Oh, enough with the false modesty — in this OP you claim to have ‘proved false’ a widely accepted body of evidence in favor of common descent.
When asked to clarify your surprising result, instead of taking the opportunity to engage, you repeatedly try to change the subject, then retreat to tone trolling.
That’s not a good look.
You also have some pretty strange misconceptions regarding the importance of basic background information in understanding any branch of science. You may be a victim of your educational background.
Nonlin.org,
Define a ‘kind’, then, and provide examples and criteria for the distinction. Bill Cole says they are hard to define. Prove him wrong.
eta – Posted before seeing Corneel ask much the same.