Here, one of my brilliant MD PhD students and I study one of the “information” arguments against evolution. What do you think of our study?
I recently put this preprint in biorxiv. To be clear, this study is not yet peer-reviewed, and I do not want anyone to miss this point. This is an “experiment” too. I’m curious to see if these types of studies are publishable. If they are, you might see more from me. Currently it is under review at a very good journal. So it might actually turn the corner and get out there. An a parallel question: do you think this type of work should be published?
I’m curious what the community thinks. I hope it is clear enough for non-experts to follow too. We went to great lengths to make the source code for the simulations available in an easy to read and annotated format. My hope is that a college level student could follow the details. And even if you can’t, you can weigh in on if the scientific community should publish this type of work.
Functional Information and Evolution
http://www.biorxiv.org/content/early/2017/03/06/114132
“Functional Information”—estimated from the mutual information of protein sequence alignments—has been proposed as a reliable way of estimating the number of proteins with a specified function and the consequent difficulty of evolving a new function. The fantastic rarity of functional proteins computed by this approach emboldens some to argue that evolution is impossible. Random searches, it seems, would have no hope of finding new functions. Here, we use simulations to demonstrate that sequence alignments are a poor estimate of functional information. The mutual information of sequence alignments fantastically underestimates of the true number of functional proteins. In addition to functional constraints, mutual information is also strongly influenced by a family’s history, mutational bias, and selection. Regardless, even if functional information could be reliably calculated, it tells us nothing about the difficulty of evolving new functions, because it does not estimate the distance between a new function and existing functions. Moreover, the pervasive observation of multifunctional proteins suggests that functions are actually very close to one another and abundant. Multifunctional proteins would be impossible if the FI argument against evolution were true.

stcordova,
This is actually nicely illustrative of a point. Why on earth would a good scientist NOT consider possible objections to their theory? When, conversely, do you?
colewd,
It might help, at this juncture, if you say what you think Dr Swamidass’s claim actually is, and why ORFan genes oppose it.
stcordova,
I read you as agreeing on protein structure, and the lability of individual sites within a conserved structure, not ‘the eye’.
Sal,
Does this mean that you accept common descent for sequence conservation, but reject it for structural conservation with complete loss of alignment?
Allan Miller,
This is his claim. Perhaps Dr Swamidass can explain how he intends to support the above hypothesis.
Threads like this always somehow remind me of the (apocryphal) “proof” that the bumblebee cannot fly. Devout believers in magicflightism have a strong need to support their conviction that, absent divine intervention, bumblebee flight would indeed be impossible.
What is presented is either “common sense” arguments (such as, just LOOK at that fat body, those tiny wings, how thin the air is. Prima facie impossible without divine assistance). Or else complex mathematical calculations carefully constructed to “prove” that natural flight cannot be performed given the constraints and limitations of the bee’s physiology.
Both sides agree that the bumblebee DOES fly, of course. Both sides apparently agree that IF there’s such a thing as otherwise undetectable divine assistance, then it can’t be ruled out. The debate is over whether such assistance is required. This debate cannot be resolved.
The are 4 levels of protein structure:
1. sequence (primary structure)
2. secondary structure (like alpha helicies, beta sheets), the “lego” parts
3. tertiary structure, the complete 3d structure
4. quantenary structure (number and arrangement of multiple folded proteins subunits).
Similarity of secondary and teriary structures does not imply common descent whereas similarity of primary structure might. In fact, here is a list of known proteins with the same tertiary structure but not the same evolutionary origin:
http://scoppi.biotec.tu-dresden.de/abac/
colewd,
This does not say that every protein, ever, is obtained by amendment of an existing one, and all such relationships should be detectable else my theory would absolutely break down …
I am not arguing this at all.
I did not say that “God cannot do this.” What an absurd claim. Obviously He can do whatever He wants, including creating life through an evolutionary process, and specially creating life in a way that looks like evolution. He does not answer to us.
And I am making the simple claim that mindless processes can produce sequences with high MISA. That is it. This is just a simple statement that 1+1=2, not that 1+1=4 as is misreported in the literature.
I have good will to Durston. I have no personal problem with him. I’m just correcting his math.
Please show me where he says that “evolution can easily generate sequences with high FI, FSC, or MISA with neutral drift.” Please present evidence, the form of specific quotes, that I have misrepresented him. I have actually presented quotes that he thinks he has “definitively falsified” evolution. Until you show me otherwise, from actual evidence, it is hard not to receive accusations like this as slander.
Thank you Rumraket for correctly representing my point.
stcordova,
Since the same 3D structure can be achieved by any number of different amino acid sequences, absence of primary structure alignment is no strike against common descent for common structures. It depends on the richness of your dataset as to whether common origin can be reliably inferred. But it cannot be reliably dismissed without a similarly rich dataset.
stcordova,
That’s not a list of proteins with the same tertiary structure.
ORFan and de novo genes are an entertaining and important, but not salient to the current conversation. Throwing this into the debate (not blaming you Alan) is a great example misdirection.
We can talk about ORFans in an article in the future. But this is only relevant to this current discussion in 1 way. We have clear evidence that some ORFans are functional and arose from untranslated DNA (e.g. nylonase), with only a few number of mutational steps. This is strong, independent evidence that functional folded proteins are not hard to evolve. It is a clue that there something wrong with Durston’s math, which should make it no surprise that I have demonstrated there is a problem with his math.
Now as for the details of ORFans and de novo genes, I suggest we save that for another thread, where I would gladly interact with Nelson and Bugg’s work. (Bugg’s by the way is a very accomplished plant biologist in the UK who recently got a very nice paper in Nature). The key point though is that if you look for ORFans between humans and chimps, there are none. We do not need ORFans to understand that piece of evolution. Which is an important point that needs to be emphasized here. I will share more in the future, but for this thread, I think the focused emphasis is on FI, FSC, and MISA.
This part I agree with. At a basic level it is correct to say that just because they are similar, does not mean they originate from a common ancestor.
That means that, in a case where all you have to go by is mere similarity, but you don’t have sequence convergence to bridge the gap and therefore no tree topology, you cannot conclude common descent. In such a case biologists will honestly and openly state that they cannot distinguish convergence from common descent. In other words, are they similar because they share common ancestry, or are they similar because of some common adaptive constraint (and that could be natural selection, or yes it could even theoretically be because it was designed to “fit the purpose”).
But it is often the case that we DO have tree-structure, even for highly dissimilar sequences. We have sequences that “bridge the gap” so to speak. So at one “end” at the phylogenetic tree, on a branch there we have a sequence that is almost entirely dissimilar to a sequence on “the other end” of the tree. And if those were the only two sequences we had, even if they folded into the same structure, we could not infer common descent. At best we could just say they were convergent, but we would not know WHY they were convergent in structure (though biologists will often stipulate natural selection in that case).
But as I said, we usually have sequences that “bridge the gap”. Next to the branch on one “end” we have one that is sliiightly different. And next to that one, one that is even more different. And so on and so forth. Through the totality of those sequences we can see a progression from one to the other. Not that they evolved by that route, one directly into the other, but the data, in terms of how the sequences nest into groups, and how similarity drops off further and futher away, we can show that they share descent, despite their almost total dissimilarity, at the most mutually distant branches.
So when PFAM says Insulin, Relaxin and others are in the same protein “family”, despite total sequence dissimilarity between some members, it is because when all the sequences are considered together, they yield an obvious tree, with sequences that bridge the gaps between the most dissimilar members.
But you knew all this Sal.
Exactly.
It is so often commonly misunderstood by creationists. Similarity implies common ancestry. No, it doesn’t, no biology thinks it does. NESTING PATTERNS of similarity do. And we have those, we don’t just have merely similar sequences.
swamidass,
I didn’t mean to suggest that you were doing something inappropriate. I believe that your paper should give a pretty good idea of how to replicate your computational experiments, and that the Python code should be strictly a supplement. Code is not an appropriate presentation of an algorithm.
My original code can be used to replicate the first simulation experiment. In the revised code, each amino acid is mutated (nominally) with probability
where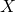 is a Poisson random variable with parameter
is a Poisson random variable with parameter 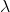 (called
(called 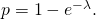 Nominal mutation of an amino acid results in mutation with probability
Nominal mutation of an amino acid results in mutation with probability 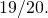 and the actual mutation rate is
and the actual mutation rate is
mutation_ratein the code). The thresholding operation essentially turns the Poisson random variable into a Bernoulli random variable with parameterIn concrete terms, when I equate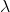 with rates given in Section 2.1, and feed
with rates given in Section 2.1, and feed 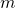 into my original code as the mutation rate, I obtain the reported results.
into my original code as the mutation rate, I obtain the reported results.
My principal concern is not the code, but instead the correctness of the results. With mutation rate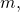 the empirical distribution for each site in the sequence of amino acids, as the sample size goes to infinity, is
the empirical distribution for each site in the sequence of amino acids, as the sample size goes to infinity, is 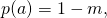 where
where  is the ancestral amino acid, and
is the ancestral amino acid, and 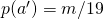 for all
for all 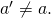 Then the entropy of the asymptotic distribution is
Then the entropy of the asymptotic distribution is
where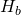 is the binary entropy function. The MISA for the site, as the sample size goes to infinity, is the maximum possible entropy minus the entropy
is the binary entropy function. The MISA for the site, as the sample size goes to infinity, is the maximum possible entropy minus the entropy 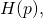
Given that sites of the ancestral protein are mutated independently of one another, the asymptotic MISA for a sample of proteins of length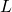 is simply
is simply 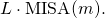 Note that I did this analysis before I did any programming.
Note that I did this analysis before I did any programming.
In the attached figure, corresponding to Figure 2 in the paper, I have drawn a line from point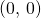 to point
to point 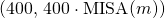 for each
for each 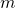 in
in
The circles in the plot are not exactly on the lines. They are data points obtained by running my original code (sample size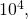 not
not 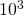 as in your code), with
as in your code), with 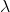 transformed into mutation rate
transformed into mutation rate 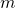 as described above.
as described above.
Dr. Swamidass:
In opening of Section 2.2, you write:
I can read “several” only as “three,” and for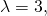 mutation rate
mutation rate 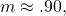 and the MISA of the sampling distribution for a length-150 protein is about 4 bits. Assuming a fix of the mutation bug, the relation of mutation rate
and the MISA of the sampling distribution for a length-150 protein is about 4 bits. Assuming a fix of the mutation bug, the relation of mutation rate 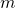 and parameter
and parameter 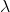 is
is
The mutation rate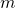 approaches 1 as
approaches 1 as 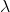 goes to infinity. Plugging in
goes to infinity. Plugging in 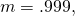 I obtain
I obtain 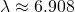 (with a note that I’m tired today, and hence prone to silly little errors). You should not suggest that there’s a minimum in MISA when
(with a note that I’m tired today, and hence prone to silly little errors). You should not suggest that there’s a minimum in MISA when 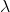 matches the number of bases in a codon.
matches the number of bases in a codon.
I am not a biologist. But I think that a biologist, computational or otherwise, must regard it as an error when “mutation” is a mutation with probability 19/20. The easiest fix for your code is to set all amino acids to 0 in the ancestral protein, and to draw mutants uniformly at random from {1, 2, … 19}. In the initialization method of
UniformSimulation, changeself.ANCESTRAL = self.random()to
self.ANCESTRAL = np.zeros(20)In the
mutatemethod, change the 0 inR = scipy.stats.randint(0, 20).rvs(self.L)to 1. I don’t have a copy of your code to run (is the notebook, rather than the PDF, available somewhere?), so I’m only 92.57329 percent sure that making the ancestral protein a string of 0’s causes you no problem.
If I were you, I’d also change the Poisson distribution to Bernoulli, and revise Section 2.1. It’s not much work, and it doesn’t have a significant impact on what you’re trying to convey. The benefit is that it’s much easier to give a simple and accurate description of what you’re doing. In the initialization method, change
self.POISSON = scipy.stats.poisson(mutrate)to
self.BERNOULLI = scipy.stats.bernoulli(mutrate)In
mutatechangeS = self.POISSON.rvs(self.L)to
S = self.BERNOULLI.rvs(self.L)Something like that, I sorta kinda almost guess.
Tom, to swamidass:
Tom,
That won’t work, because after the first mutation at a given position, subsequent “mutations” can still be non-mutations in your scheme. For example, suppose position 5 has mutated from 0 to 7. When it mutates again, it’s possible for the random number to be chosen as 7, in which case the new mutation isn’t a mutation at all.
Another flaw with your scheme is that you are preventing any reversions to the ancestral state at a given position. That is, none of the amino acids can ever revert to zero.
My suggestion would be, when mutating, to draw a random number from {1, 2, … 19}. Make the new amino acid equal to the sum of the random number and the current amino acid, modulo 20.
Then:
1) the mutation rate is accurate — that is, all mutations involve actual amino acid changes;
2) the ancestral sequence can be initialized randomly; it doesn’t need to be set to all zeroes; and
3) reversions to the ancestral state are possible at a given position.
keiths,
Read the paper carefully, and you’ll see that all “extant sequences” are obtained by mutation of a single ancestral protein. If you remain unconvinced, well, it takes only a couple minutes to read the relevant portion of the Matlock-Swamidass code.
Tom, I appreciate the effort here. If you email me I can send you the python notebook. My email is easy to find online. If you can clean this up, perhaps we could make a public code repository to let other people run these simulations themselves. Just a thought.
Thanks for looking at this and the points you raise.
First off I find it entertaining that the only one looking at the actual paper and the code (and even finding some rough spots) is someone who agrees with the conclusions. I wonder why that is? The best way to counter this paper is to find an error.
Second, the points you are raising are confined exclusively to the “uniform” mutation rate and are in reference to simplification we used to improve simulation efficiency. Your point is taken that we should mention this in the paper, but there is no error. It is common use “sampling with replacement” to model processes that are actually “sampling without replacement”. It turns out that they very quickly converge to the same solution https://www.ma.utexas.edu/users/parker/sampling/repl.htm; but sampling wiht replacement is much more analytically tractable. That is what we are doing here, and one could easily change the code but this will only affect to final results a tiny amount (probably less than 1%). I think this approximation is what is confusing you, and it is not an error.
I am not using mutation rate to reference the number of differences between the ancestral and extant sequences (i.e. the extant divergence). Rather, is the number of mutation events per amino acid, but averaged across the whole protein. So some amino acids may be mutated more than once. It might be more clear to say this is the “mutation event rate”, rather than the “mutation rate”. But this is also a common way to do forward simulations because it allows explicitly for “reversion to ancestral” mutations.
Another trick that enters this model is that sampling with replacement one time produces the exact same distribution as sampling with replacement multiple times (this is not true for without replacement sampling). So we take a shortcut and just sample one time if the number of mutations returned by the poisson > 0. So if you want to move to sampling without replacement, you have to mutate multiple times based on the output of the poisson distribution. So your fix to my code will actually break it, because you are not doing the mutation without replacement multiple times as you should.
And it is correctly a poisson distribution, it is not a bernoulli distribution. The output of the poisson distribution is the number of times a mutation occurs at that amino acid, even if it eventually ends up at the starting amino acid.
Remember, that the asymptotic distribution for both with replacement and without replacement mutations is the same.
In your math you are computing the expected MISA for your “fixed” model, but you are not mutating multiple times as you should if you use the without replacement model. A correct analysis would show the asymptotic MISA as lambda increases is 0, and you get there very slightly quicker with my model. Also “several” and “approaches” is deliberately vague, because the exact speed of convergence is beside the point. The ranges I used here cover the equivalent more than 100 million years of mammalian evolution, and over this range we still see a very strong effect of “shared history” in the simulation.
In the end, none of these points are important to the final results. As you can see, you reproduce the graph almost perfectly. I agree I need be more clear in the manuscript (and will make changes accordingly), but this only applies to the uniforum simulation (the codon simulator does sampling without replacement), and does not change the results or the conclusion.
Once again, Tom, thanks for looking at this. Your points are going to shape the next draft, so I can be more clear about this in the writing, and remove any confusion. Of course, those that do not think these are valid approximations, can always do a more exact simulation (and release their code for review). They will find nearly exactly the same results.
Not meaning to weigh in on a tussle between friends…
This is correct. But I will emphasize that this does not change the results appreciably from my code. Almost not at all.
Tom’s fix though will produce a slightly different result at the asymptote. His simulations MISA will got to log 20 – log 19, not zero, as lambda goes to infinity.
Never? I thought these high level languages were supposed to change that. 🙂
Why wouldn’t I use his own words? To be quite frank, anyone who reads Durston’s paper can figure it out for themselves. That would include you, Rumraket.
But because i like you, let’s have a look. 🙂
Durston:
So in my own words, a Darwinian evolutionary process. And by impossible, it cannot be expected to happen even once in the history of the universe.
So not impossible as can never happen, as swamidass claimed in the abstract, but rather so highly unlikely as to be practically impossible.
Durston again:
That’s the kind of evolution he’s talking about.
As a theistic evolutionist Swamidass holds that God is required for evolution. There are no “Godless” evolutionary processes. If it were otherwise, “theistic evolution” would be an oxymoron.
If Swamidass holds that some evolution is atheistic but other evolution is theistic then he is no different from the IDists that he pretends to be arguing against.
What does it even mean to say that God could create life by an evolutionary process? Do you mean “chemical evolution”?
Because if your math and your simulation is about the origin of life rather than it’s diversification from one or more common ancestors, I missed that.
swamidass,
We agree that Tom’s “fix” is broken, but he was right to point out your 19/20 mutation rate error.
My fix addresses both of those problems.
As for this…
…I would argue that it’s better to have an accurate model than an inaccurate one, especially when the fix is so easy. If you use an inaccurate model, you have to demonstrate (at least to yourself, and ideally to the reader) that the inaccuracies don’t matter to your conclusions. No such problem with an accurate model.
Also, you might end up reusing this code for future work, or someone else might adapt it for their own purposes. Better for it to be accurate in those cases.
The fix seems worth it for all these reasons, especially since it is so easy.
And you’re going about this by using an intelligently designed “simulation”?
Good grief, Mung. You’ve been at this for 15+ years, and you still don’t recognize how stupid that argument is?
Didn’t you say that your student wrote the code? There aren’t many ways to read
def extant_sequence(self):
return self.mutate(self.ANCESTRAL)
which is called from
def run_simulation(m, L, iters, simcls):
S=simcls(L=L, mutrate=m)
for i in xrange(0, iters):
nseq = S.extant_sequence()
ndiff = S.diff(S.ancestral(), nseq)
S.accumulate_sequence(nseq)
yield (i+1, ndiff, S.MISA())
Every new sequence
nseqis obtained by mutation of a single ancestral sequence, set heredef __init__(self, L = 300, mutrate = .5):
super(UniformSimulation, self).__init__(L)
# Initialize a random ancestral sequence.
self.ANCESTRAL = self.random()
# Initialize a poisson mutation model
self.POISSON = scipy.stats.poisson(mutrate)
when the
UniformSimulationclass is instantiated. The use of uppercase in the nameANCESTRALmeans “TREAT THIS AS A CONSTANT.” I just verified that the only assignments toANCESTRALare in the initialization methods of the two subclasses ofSimulation.As I wrote above, all “extant sequences” are generated by mutation of a single ancestral protein — not only have I matched your results by doing that in my code, but I’ve shown it to you in your own (student’s) code. If the ancestral protein is set to a sequence of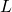 0’s, instead of randomly, then mutant amino acids are uniform on {1, 2, … 19}. That’s what i’ve done in the code I used to generate close matches to your Figures 1 and 2. And if I were wrong about that, my results would NOT be close to yours. Of course, this does NOT work for
0’s, instead of randomly, then mutant amino acids are uniform on {1, 2, … 19}. That’s what i’ve done in the code I used to generate close matches to your Figures 1 and 2. And if I were wrong about that, my results would NOT be close to yours. Of course, this does NOT work for
CodonsSimulation.What you’re doing is important, and thus is important to get right. I’m taking great care to give you accurate feedback. (I’ll say more about your interpretation of the paper by Durston et al. when I complete the painful process of reading it.) I ask only that you process my feedback carefully. I understand that you’ve got people coming at you from all directions, and realize that you may have lost track of which simulation I’m addressing.
I do appreciate the help. Let me request that we do this offline in an email chain. That way I can include my student and keep track of things. At this point, I cannot commit to answering all your questions on the website immediately, and I do not anyone to be left with the false impression that I made fundamental error that changes the results. To be clear, I would publish a retraction if a serious error is actually uncovered. But this type of critical feedback is too valuable to exchange in blog comments. Like I said, you can find my email at my website: http://swami.wustl.edu/contact.
I agree and this is the right solution.
Yes I know. But that is because (as I have already explained) this is not the right simulation. You need to repeatedly mutate the sequence for each count returned by the poisson distribution. You also are mistaking “mutation rate” for “divergence (the number of differences)”. With those fixes, it will come back to the right answer.
This is an intelligently designed simulation of a mindless process. This demonstrates the mindless process can produce sequences with high MISA.
The impression I got from your paper was consistent with what you’ve just written. But that’s not your (student’s) code actually does:
def mutate(self, seq):
# Sample mutation events from poisson distribution
S = self.POISSON.rvs(self.L)
# Random vector of AA selected uniformly from AA
R = scipy.stats.randint(0, 20).rvs(self.L)
# Apply mutations to sites
return np.array([(r if s>0 else aa) \
for aa, r, s in zip(seq, R, S)])
The code and “success” otherwise). Again, if this were not the case, my original code would not produce your results.
and “success” otherwise). Again, if this were not the case, my original code would not produce your results.
for aa, r, s in zip(seq, R, S)])iterates over amino acidsaain a sequence, along with possible replacementsr(drawn uniformly from {0, 1, … 19}) and Poisson variatess. The coder if s>0 else aareplacesaawithrif and only the Poisson random variatesis greater than 0. When the Poisson variate is dichotomized in this fashion, it is actually a Bernoulli variate (“failure” whensis 0, i.e., with probabilityWhen I bashed out my little code, I did not think I was implementing your algorithm. I planned to use it for comparison. That’s why I wrote “for what it’s worth” when I posted it, along with a plot analogous to your Figure 1. I was surprised when I saw that your code actually does the same thing as mine (for a different mutation rate).
It appears that you expected your student to do something other than he did. (I’ve had that experience a time or two or three.) I’m assuming that you prefer to learn this before publication, rather than after. Please tell me that the paper has not been accepted already.
I’ve missed that explanation. You’re saying that you know that the code in the notebook is wrong?
The student’s code is consistent with what I wrote. (1) we use a poisson distribution (P) with a rate of m to compute the # of mutations at each site. (2) we use the approximation of sampling with replacement for each mutation. (3) under this approximation, mutating a site one time produces the same distribution as mutating it multiple times, so we only need to mutate one time if P>0, or none if P==0. This is exactly what those three lines do.
This is nearly equivalent to sequentially applying single mutations to the sequence, and actually converges a little quicker to 0 MISA. It is a common approximation in forward simulations that improves efficiency.
The notebook code is not wrong. Your fix, the one that produces a different graph, ends up being wrong.
If you insist that we use mutation without replacement (not using approximation #2 above), then you cannot use trick #3. You have to, instead, mutate each residue number of times returned by the Poisson distribution. If you do this, you will see the results are nearly identical.
Keith proposed doing this by adding a random int from [1 to 19] and taking modulus 20, once for every mutation required by the Poisson distribution. That would work.
Mung,
Durston takes a subset – commonly descended and tuned proteins with a common function – and determines from that subset that there is no path of folded proteins linking it to other such subsets. That doesn’t work.
It could be true, but its truth isn’t established by that data.
He does no such thing Mung. In the studies I quote, Kirk does not consider pathways at all. All he considers is the number of functional sequences, not their distance to other proteins. Before you argue this obvious point, produce quotes from his paper and specific references.
The one time he does consider a pathway in the literature (you quoted it earlier too), he then computes that a new function arose with minimal information input (correct!). Then, in a non-sequitur, he concludes this example has no relevance to his prior work because no “statistically significant FI” was input into the system. Of course, this does falsify his theory because it demonstrates (using his own math) that only a tiny amount of information is required to produce a new function, a totally different number than he computes using FSC (i.e. MISA).
So my counter to him is, (1) What is statistically significant FI (SSFI) ? it is never mathematically defined. (2) we agree in at least one case that SSFI is not needed to produce a new function, what evidence is there that any function requires SSFI? (3) And does not the fact that FSC computes (as he computes it from HMMs) a very high value in this example, demonstrate it is an unreliable measure of evolvability?
I think the answers to all these questions are obvious. (1) not clearly defined. (2) there is no evidence SSFI (whatever it is) is required for new functions. (3) clearly FSC is a very poor estimate of evolvability (both because FI has nothing to do with evolvability and MISA has nothing to do with FI).
swamidass,
That’s what he appears to be doing in the paper I found by pasting Mung’s abstract into Google.
Considering pathways?
Can you point us to some undesigned models of, erm, anything.
This post was brought to you by tautology.
No doubt that’s what Durston means by impossible. To say that Durston claims evolution is impossible is to distort his position [not that keiths cares] unless it’s made clear in what sense he is using the term. Counting on an equivocation isn’t really a responsible way to write a technical paper.
What have I said that contradicts this? This is exactly what I meant by “impossible”. And you have yet to quote Durston. Looks like I have characterized his claim correctly. He claims to have “falsified” evolution with his FI argument. It is the other way around, I have falsified his argument.
swamidass,
This is standard operating procedure for Mung. He doesn’t understand Durston’s paper or your response to it, so he has nothing to contribute to the discussion. He feels left out, so he’s fishing around for a ‘gotcha’ — and failing.*
Sit this one out, Mung. It’s way above your pay grade.
* Or making inane arguments like this:
AIG keeps a list of arguments so stupid that creationists shouldn’t use them, to save embarrassment. And the “it’s not really evolution because simulation was Designed” is in that category.
Mung doesn’t give a shit about science, he just wants to talk to people.
AhmedKiaan:
A recent and very telling comment from Mung:
petrushka,
No, declaring their absence. The very quote (of Durston) which led to the paper I was talking about, says “The genesis of novel protein families must proceed via a blind, unguided random walk across non-‐folding sequence space.”. Which is to say, as I read it, there is no pathway of folding proteins linking two families of folding proteins.
Yes. Basically Durston seems to be saying that, in so far as a sequence from that area of sequence-space isn’t found in extant life, it’s because it’s a non-folding and/or non-functional portion of sequence space.
Basically that, all the sequences we see in life are the only functional ones possible, and all other sequences are nonfunctional. In support of this he uses… only the sequences from extant life. In other words, circular reasoning. He can’t extract that conclusion from the data. He can not extrapolate from used sequences into unused sequences and say the reason they’re unused is that they’re nonfunctional and that evolution would have to wander aimlessly around in this vast sea of nonfunctional polymers and by some grand miracle of luck, happen to find another of the functional ones.
We discussed all this with Kirk who was nice enough to drop by and interact in the comments, back here.
I read that to mean he has considered pathways and denied that they exist. A claim that is incompatible with Wagner’s. If Wagner is correct, Durston is complete rubbish.
Which might explain the resistance to Wagner.
This would appear to be a Thunderdome. Two go in, one comes out.
Leaving aside the lack of credibility of the people you quote, the environment is a great source of information about what works and what doesn’t.