I see long-time commenter at Uncommon Descent, Mung, in a thread entitled Backwards eye wiring? Lee Spetner comments, asks:
How do you calculate the size of amino acid sequence space?
As this seems somewhat off-topic there, I thought I’d attempt to answer Mung’s question. I’ll try and be brief. The two most fascinating biochemicals are nucleic acids (RNA and DNA) and proteins. Proteins seem ubiquitous in cellular systems; they function as catalysts (enzymes), structural elements (keratin, collagen), signal molecules (hormones, pheromones), binding agents (antibodies). Proteins are linear sequences of amino acids joined by a condensation (called so because a molecule of water is lost) reaction forming a peptide bond. There are twenty-one amino-acids found in eukaryotes and twenty of them are directly represented in the genetic code. The special case is selenocysteine which is coded indirectly and I’ll leave that out of the calculation for the sake of simplicity.
So what number of different amino acid sequences could theoretically exist, given twenty possibilities for each aa in the polymer. I guess we shouldn’t count twenty monomers. For dimers, there are 400 possibilities. For trimers, we have have 8,000 and so on. The general formula for the number of theoretically possible different protein sequences of length ![]() is
is ![]() . So the answer for all possible sequences is the sum of this calculation from
. So the answer for all possible sequences is the sum of this calculation from ![]() to, well, what? There are some very large proteins; titin being the largest known at around 30,000 aa’s. So I guess we should sum at least to that number.
to, well, what? There are some very large proteins; titin being the largest known at around 30,000 aa’s. So I guess we should sum at least to that number.
This is a very big number indeed! I leave it as an exercise for the reader to try representing the number that results when taking the upper limit of ![]() as 30,000. 🙂
as 30,000. 🙂
Now I’ve answered Mung’s question, would he like to enlarge on what it signifies?
ETA categories and remove tautology
ETA 2 correction ![]() not
not ![]() (hat tip Joe Felsenstein)
(hat tip Joe Felsenstein)

You’re interactions here with me here could be improved if you learn to read for context and drop the silly insults. I don’t think any such thing as what you imply. I was responding to a post someone else made about folds and said I could go simple folding one better.
There are in fact a class of proteins where folding is only part of the story, and that was my point. Not that all proteins are enzymes. Sheesh.
Yes, I understand that you’re wrong and that you made the same mistake that Nick did. The size of the space is 20^n. You don’t reduce the size of the space by introducing more functional proteins into it.
Well. That leaves me out. Can I try anyways?
Let’s take two amino acids with sequence length of 2 = 2^2 = 4. I bet everyone can get that one right. 2 with length 3 = 2^3 = 8. 4 with length three = 4^3 = 64. Oh wait, that last one is nucleotides and codons.
Sequence space. You can decrease the size by reducing the number of letters in the alphabet or reducing the length of the sequence. What other way is there of reducing the size of the sequence space?
Allan:
Preach it brother.
Mung,
So you don’t understand. As I expected.
As I wrote in the comment that you quote…….d, the size of the sequence space is utterly irrelevant.
I have an aversion to outperforming expectations. Then people come to expect that from you. It’s a vicious cycle.
If the size of the sequence space is irrelevant, why didn’t you just say so in the first place? Why spin up the numbers? What on earth does the size of the sequence space have to do with how easy it is to navigate? Sheesh.
That is the very question we’ve been asking IDists for years.
Mung:
petrushka:
Indeed. Mung, why don’t you ask KF that question the next time he trots out the “islands of function” argument?
Mung,
I think there is some confusion (on both sides to some extent).
The absolute size of protein sequence space has no direct impact on the amount of function it contains, nor its connectedness, as you say. This cuts both ways, and effectively undermines Hoyle, because that’s what he bases his argument on. He assumes a single target of length n in a 20^n sea. That is clearly bogus. He just ignores connectedness.
Simply because of redundancy at the amino acid level – a given site can accept one of a number of acids of similar properties at the majority of points on a given protein – the ‘single target’ model is clearly wrong. If we imagine colouring in the wider space, marking functional proteins in red, we evidently do not have a single red dot, but an entire collection of red dots clustered around a neighbourhood. And that collection is actually going to be pretty big. And because it’s clustered, it is explorable. This does not necessarily get us from one function to another, but it does allow tuning of a lucky hit.
One could represent it another way – rather than colouring in the 20^n space, we could say that the space is actually based on residue property, not identity. If, for example, a given site will accept any polar residue, it doesn’t contribute 20 to the total space size, but a smaller number depending on how polar/nonpolar residues are distributed in the 20-acid alphabet. This is a more complex calculation, and pretty much impossible to do accurately without empirical investigation due to the subtle effect of every substitution on the rest of the chain.
Nonetheless, that is Nick’s point, and the point of the paper he linked – because of extensive redundancy, the actual space of different proteins – those whose properties, rather than raw sequence, are different – is much smaller than that assumed by the ‘digital’ approach.
One could analogise to a language where a single word had exactly the same meaning acrioss a range of apparent ‘misspellings’ – where banana, bernarner, badada, bynana etc were all functionally equivalent. One could either say they were multiple dots in a 26^6 space, or that there was a single ‘functional’ dot in a smaller space that took account of substitutalbility.
The latter is pretty much what is done in generating libraries of random peptides. You know that only a fraction of the entirety of space will fold, so you create a patterning algorithm – say PNPPPNNNPNPP where P is polar and N is nonpolar. If there are 7 Ps and 6 Ns (say), the space is 7 * 6 * 7 * 7 * 7 * 6 * 6 * 6 * 7 * 6 * 7 * 7 = c64 million, not 20^12 – a c640,000-fold difference. This algorithm generates only a fold, not a particular function – it merely saves wasting time in potentially ‘dead space’. If you generate a million random peptides using the algorithm, you have explored 1/64th of that space.
But this would not be worth doing at all if the remaining subspace were itself largely devoid of function. This takes us back to navigability – the ‘islands of function’ argument. In the example, peptides fitting the algorithm are a mere 1/640,000th of the space of 12-acid peptides. If we assumed for argument that all peptides had to follow that algorithm, only 1.5 millionths of the 20-acid space is available to explore. But within that 1.5millionths of the space, there is clearly very high interconnectedness. Any P can be substituted by 6 others, any N by 5. Of course the result may not be functional. But that is not a result of the fact that there are 20^12 different 12-acid sequences – the vastness of the unexplorable part of the space is completely irrelevant to the exploration of the part where function is found (in this toy example). Therefore, it is bogus to invoke it.
One could try answering 3 simple questions. If one had a primitive system consisting of 4 amino acids, and added a 5th to it, the total space of a 10-acid peptide goes from 4^10 to 5^10 – c 1 million to c9.7 million. Let’s say each acid has the same chance of being in a functional protein as any other.
Would the percentage of the functional portion of the entire space be expected to
a) Rise
b) diminish
c) stay the same?
Would the interconnectedness of the entire space be expected to
Would the percentage of the functional portion of the space be expected to
a) Rise
b) diminish
c) stay the same?
Would the interconnectedness of the space neighbouring existing functional peptides be expected to
a) Rise
b) diminish
c) stay the same?
Why do the calculations? Because you asked.
You may not be aware of this, but many IDists base their improbability arguments on the size of the sequence space.
Allan Miller,
This was the point I was trying to make with my “thought experiment for any honest IDist”. Thank you for making it far more explicitly (and with less snark) than I ever could.
Now we sit back and wait, I guess…
That’s great Mung, then what the hell use is it to insist that the size of sequence space is 20^n when that number isn’t at all informative about protein evolution?
I have read for context and not used any insults. I think our interactions could be improved if you dropped the faux offense and tried to appreciate that as an outsider to the initial conversation you had with Nick Matzke, I was not aware of what particular subject you were discussing, I merely read the post of his that you yourself linked and tried to glean the context from that.
I already affirmed that the size of the space is 20^n. I have made no such mistake, in fact I agreed that when Matzke initially seemed to deny this he was making a mistake.
You would have understood this if you read the part of my post where I write that “He’s clearly made a minor mistake there when he says sequence space isn’t 20^n. The size of sequence space is 20^n for a 20 amino acid alphabet.” – and I explained why. As such, your reply here is baffling.
No, you don’t reduce the total size of the space, but you reduce the size of the space you have to search before you hit on something functional.
On average, you will have to search less (do fewer iterations, spend less time iterating) if a larger fraction of sequence space is functional, before you get *hits*.
The total size of sequence space remains 20^n, but when it comes to searches for “something functional” it can in practice be considered a space of (>20)^n.
DNA_Jock,
Cheers. Shame I couldn’t catch the bloody copy-paste transposon before the edit window ended though! I hate when that happens. 🙂
Allan Miller,
2nd question should read
Would the interconnectedness of the entire space be expected to
Would the percentage of the functional portion of the space be expected toa) Rise
b) diminish
c) stay the same?
Now Mung will have a field day over my tendency to err – curses!
What would you like me to ask him, why he doesn’t throw more islands at the problem?
Because the size of amino acid sequence space is 20^n.
No one here asked Alan Fox to trot over to UD and take a quote of mine out of all context and try to make hay out of it.
Contrary to Nick’s post and run claim, calculating the size of amino acid sequence space by taking 20^n is not “bogus.” It’s a perfectly legitimate way to calculate amino acid sequence space, and I don’t know of any other way to calculate amino acid sequence space.
Like I told Nick over at UD, you can reduce the size of he space by pretending there are fewer than 20 amino acids.
The theoretical sequence space (assuming 20 aas) is bigger than that. It is the sum of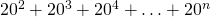 where
where 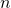 can be at least 30,000. The trap Axe, Durston and KF (with his ludicrous islands analogy) fall into is claiming unknown proteins lack function whereas the most that can be said currently about unknown proteins is that we don’t know if they might have function in some scenario.
can be at least 30,000. The trap Axe, Durston and KF (with his ludicrous islands analogy) fall into is claiming unknown proteins lack function whereas the most that can be said currently about unknown proteins is that we don’t know if they might have function in some scenario.
Oh come now! You appeared to want an answer to a question. I provided it. Other contributors – I have to say Allan Miller has been sterling in this regard – have chipped in to provide you with a wealth of information. If you don’t want to know, don’t ask!
You don’t reduce the size of the space by throwing more protein folds into it. There are two ways to reduce the size of amino acid sequence space. 1. Reduce the length of the sequence. 2. Reduce the number of amino acids.
You don’t have to search the whole space if functional proteins are not isolated in vast seas of non-functional sequences. You just have to find something that works. Then you can fine tune it.
… and? Is that the only thing you set out to highlight? The size of amino acid sequence space is 20^n. Just a blanket statement of an uninformative factoid?
So all you wanted to do was calculate the size of amino acid sequence space, that’s it? All you wanted was a simple answer to a simple question. Well then I guess we’re done.
I agree. The total size of the space remains 20^n no matter what. The size of the space is 20^n.
Where do we go from here?
You only have to try nearby variations of what you have. We live on the surface of a planet. Humans covered the surface by walking. They did not have to test every cubic meter of the planet to find the habitable spaces. They found them by exploring what was nearby.
Appearances can be deceiving. If you’re going to go grab something I wrote over at UD and turn it into an OP over here at TSZ you might want to consider the background to what was written over at UD, else you’re not doing anything much different from just quote-mining.
Absolutely nothing. Which is what happens when you take it out of context. It was a response to a hit and run post by Nick Matzke at UD with a history and context at UD that was not reflected in the OP.
Mung,
I’m pleased you’re against ‘hit and run’ posts, Mung!
Probably nowhere. I wasn’t making any sort of argument about how prevalent functional proteins might or might not be nor was I making any argument about how easy or hard it is to navigate the sequence space.
Yes. Perhaps one of these days Lizzie will have time to do more than post and run.
Besides, you’re all a bunch of teddy bears. I would not want to hit any of you.
[Strangle maybe]
OldMung:
Thus confirming my suspicions about what happened to NewMung.
Fair enough Mung.