I often encounter posters here at TSZ who claim that Genetic Algorithms (GAs) either model or simulate evolution. They are never quite clear which it is, nor do they say what it means to model or simulate evolution (what would be required) and how GAs qualify as either one or the other. My position is that GAs neither model nor simulate evolution. In addition to other reasons I’ve given in the past I’d like to present the following argument.
GAs are often used to demonstrate “the power of cumulative selection.” Given small population sizes drift ought to dominate yet in GAs drift does not dominate. Why not?
Three questions:
- How do we determine the effective population size for a GA?
- How do we calculate the value of the selection coefficient?
- How do we determine when genetic drift will overcome the effects of selection?
In a GA written by keiths (a version of the WEASEL program) the default population size is 200.
#define POPULATION_SIZE 200 // total population size
Effective population size is the number of individuals in a population who contribute offspring to the next generation.
Even though the population size is 200, only one is selected to contribute offspring to the next generation.
#define NUM_SURVIVORS 1 // number of survivors per generation (must be less than POP_SIZE)
Given an effective population size of one, drift ought to dominate, but it doesn’t.
Given an effective population size of one, what must the selection coefficient be for drift to not dominate selection?
I’d truly appreciate any assistance with the concepts or the math.
In any event, there is no way that this GA (the keiths WEASEL program) either models or simulates evolution.
Reference:

Weasel doesn’t latch, so a favorable mutant doesn’t have to replace the original.
Usually, but not necessarily.
Flint,
Had Mung known that Ne * s was large, he would have had even less of a reason to assert that drift “ought” to dominate selection.
It’s right there in the article he linked to:
In reality, Mung didn’t know the value of s, which is why he asked in the OP:
Since Mung didn’t know the value of s, he had no basis for his assertions that drift “ought” to dominate selection.
Mung,
And you invoke it because evolution stalls for lineages that fall outside this ‘Happy Zone’? Hmmm. Of course this can only happen as a result of a Celestial Finger pushing them back, just as our planet is kept just so by the same finger. Praise the Lord.
If the parameters are indeed fine tuned, and GAs work just fine as evolution in action, then evolution itself must be capable of something. Feathers, whales and flagella can surely evolve without further prodding, and the rest of ID, beyond that initial parameter setting, is bollocks, derived from insufficient faith in the foresight of the Designer. That, or God designed a process that does not do anything much.
The perpetual unhappy zone of the IDist. Evolution is nonsense, but God designed it and it’s great.
Incidentally, the population size in Weasel isn’t really 1. It crashes to 1 every generation, but in order to have any variation at all, you need variants in the plural. It’s emergent, heh heh.
Allan:
Better to bimble about in the Happy Zone than to bumble and grumble in the IDers’ Unhappy Zone.
Nope. Weasel is very close to, but not quite the same as, a Wright-Fisher model. In that model there are not 200 offspring but an infinite number of them. Natural selection, mutation, and migration change their proportions (deterministically), and then a random N chosen from that mix of genotypes survive to adulthood.
In this case there are 200 offspring (or whatever number you set) so it gets close to the WF model. In the WF model the effective population size is N. We do not say that the effective population size is infinity in a WF model. In fact it was precisely the WF model that Sewall Wright used to investigate the effect of genetic drift on gene frequencies.
In the original post Mung said two things:
Mung asked:
My answer was a bit off. The Weasel is haploid, not diploid. So the condition for effectiveness of natural selection is actually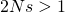 . With that emendation, I take it that Mung will agree that this question has been answered: it is
. With that emendation, I take it that Mung will agree that this question has been answered: it is 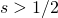 . That is a rough answer because the effect of selection gets gradually bigger and is bigger than genetic drift at about that value. I can plot curves and show this if needed.
. That is a rough answer because the effect of selection gets gradually bigger and is bigger than genetic drift at about that value. I can plot curves and show this if needed.
But Mung also said:
The keiths Weasel program is close to a Wright-Fisher model. That model is the central one for the study of the role of genetic drift in evolution. The answer to Mung’s question about genetic drift is obtained from a diffusion equation which describes the limiting behavior of a sequence of WF models as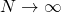 and as
and as 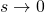 such that the product
such that the product 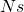 remains constant.
remains constant.
The WF model is sufficiently useful in the study of evolution, in this case the study of genetic processes in populations that it enables us to answer Mung’s question.
By the way, if anyone wants to explore the behavior of a one-locus diploid Wright-Fisher model with two alleles, and try to discover for what value of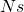 we find selection having a noticeable effect, I immodestly recommend the program PopG, a Java program that is distributed from our website, and used for teaching population genetics.
we find selection having a noticeable effect, I immodestly recommend the program PopG, a Java program that is distributed from our website, and used for teaching population genetics.
I’m confused. If the population size is N, and N is the number that survive to adulthood, then the populations size of WEASEL is one?
Yup, as a haploid effective population size.
Poor Mung’s brain is gonna explode when he reads this
That agrees with what I said, although when I said it, I was just going on intuition.
The PopG link is broken.
http://evolution.gs.washington.edu/popgen/popg.html
It is given by dazz, and I also edited my comment to fix it. Both links should work now.
In this very thread you responded to keiths thusly:
I then pointed out that your last comment on the thread where you claimed to have said “it couldn’t be done” was:
The evidence shows that both your claim to have have said “that it can’t be done” is false and your accusation of quote mining is false. A decent, honest person would therefore retract both of those claims. Is NewMung available?
Joe Felsenstein,
Sure, the effective size is 1 after a bottleneck – the ‘ideal population that would have the same variation as the real population’: Ne=N.
But it rises after births. Of course you don’t tend to look at it on such a fine scale, but I stand by what I say. If offspring are variant and potential reproducers, Ne can’t be 1 at that moment.
Flint,
I do agree with DNA Jock’s response below but in multi protein complexes and proteins that work together they form with both shape and charge matching. If you are putting together a puzzle the first piece selected can be anyone but as you move further the choices of shapes become more specific. Intuitively this would be the same for proteins.
(I typed in a long reply to this, and when I tried to post it I got a “we can’t find this page” error – and lost everything. I’ll try again – and THIS time I’ll save my work!)
I agree with you that, when you’re putting together a jigsaw puzzle, the shape of each additional piece is increasingly constrained. But this puzzle analogy is incorrect.
There is no “fixed target” in evolution. The pieces you already have can change at any time, new pieces can fit in unexpected ways, or if they can’t fit at all, they can be retained and sometimes fit later, as things change. And at the puzzle level, the nature of the entire puzzle is not fixed, or even defined.
This is why demands that evolutionary biologists predict the path of evolution (of anything) are misguided. We can understand and predict that change will occur, and that much (but not all) of it involves conserving beneficial mutations, but we can’t predict what those mutations might be, and we can’t even predict the nature of the benefit.
There’s an old story of a congress of gorillas trying to predict their evolutionary path (beneficial mutations). Would they have even considered the possibility that the “improvement” would entail the loss of fur, the loss of strength, and the loss of climbing ability? Would a congress of humans even consider the “improvement” of a loss of intelligence, or loss of language? What could we possibly gain in exchange that would be better?
colewd,
I have not yet identified anything that we do agree on. In particular, I answered your question:
with “No, I do not.”
I suggest that your intuition re proteins is in error.
Many people seem to conflate optimization and complexity. In a highly optimized system, any change is likely to be deleterious (this is pretty much a tautology). In a highly complex system, this is not necessarily the case (unless the system was designed, in which case…)
Vertebrates are complex, but NOT highly optimized. If you what to see optimized, check out bacteriophage…
It’s also why we can’t design, except by trial and error, by mimicking evolution.
Flint,
Can you show me why it is incorrect other than quoting how evolution works? Do you agree that proteins fit together because of their shapes and charge?
DNA_Jock,
Do you agree that proteins fit together based on shape and charge? If so why would the puzzle analogy not hold?
Because the “socket” where any protein must fit, is as subject to mutation as the protein that might fit it.
Again, you are assuming a fixed target. But nothing is fixed. Socket-stuffing is one of the key goals of antibiotics. Morphing the sockets is one of the key countermeasures of bacteria.
I think Joe F. explained this clearly. Complex organisms are rarely well optimized, and the number of possible optimizations at every level is simply unlimited. As a corollary, making something MORE complex increases, rather than reduces, the improvement space.
DNA_Jock,
This is quite a statement. How have you determined they are not optimized?
Flint,
With all due respect to Joe, I don’t think he has a clue what the optimization tradeoffs for life are. When we can create a protein sequence from scratch or discover how splicing is sequenced or truly understand how proteins communicate with each other to cause a cell divide then the conversation can begin of whether optimization has occurred.
Flint this does not make sense. Once the shape of protein a is formed then the mating protein no longer has freedom of sequence. Protein specific drugs work because they can conform to the shape and charge of their target.
OK, you win. Nobody knows what they’re talking about, nobody has any clues, nobody can answer your questions to your satisfaction, nobody can dislodge your convictions. I’ll leave you to ponder the mysteries of why some drugs stop working, why they have side effects that vary among people, why they work for some people but not others. Maybe you can organize your convictions and explain this to all the biologists here. They sure can’t explain anything to you.
Of course the sequences continue to mutate. Coding sequences have fewer neutral mutations available, but they nevertheless do change over time. Given enough time, the majority of bases can change.
petrushka,
I agree. Freedom does not mean 100% precision, but the shape must be compatible.
Flint,
When you can cite repeatable experimental evidence of a mechanism that causes novel gene sequences to form I will change instantly. Until then you have an untested hypothesis with lots of stretch assumptions that are not standing up to logical debate.
This whole thought is terribly muddled.
New sequences form almost with every cell replication. Each and every human has half a dozen or more mutated coding sequences.
If you mean observing a protein coding sequence mutating into a sequence for a different protein, Thornton has demonstrated how an ancestral protein can duplicate and morph into two different proteins.
If you mean completely novel sequences, we have observed random sequences that can be substituted into bacteria.
If you are looking for squirrel into bird change, no one can help you. Perhaps you could get a job in Doug Axe’s lab.
I have such a mechanism in mind. I can provide you the sequences that it outputs.
What test will you perform in order to determine that they are novel?
What do you mean by repeatable? It sounds like you are saying that if random mutations cannot be replicated, precisely and in detail, then evolution is an untested hypothesis that can’t stand up to logical debate.
You might as well argue that if a baseball game can’t be replicated down to the last detail, and thus no two games are alike, then there’s no such thing as a baseball game and anyone who THINKS they are watching baseball doesn’t know what they’re talking about.
Processes with key random elements that cannot be precisely replicated are not therefore unscientific. Casinos in Vegas have no idea what card will come up next, which pull of the handle will score on the slots, but they can provide statistics close enough that half a percent difference is indications of cheating.
Or counting.
Finally, something we do agree on!
🙂
Induced fit.
Well, I was thinking about the initiation of transcription, the initiation of translation, and inappropriate codon bias (in, say, blood-clotting factors), but if you want an example that is more lay-accessible, there’s that whole breathing-and-eating-through-the-same-narrow-tube thingie…
I understand that the sewer-through-a-recreational-area thingie is more contentious.
Only on one game. Counting slot machines won’t help you.
Why? I’m not the poor schlepper who claimed that GA’s model evolution.
“Optimize” is not a universal, one-size-fits-all process of improvement. You “optimize” something for a particular purpose, and the results of that “optimization” can be very different indeed, depending on the specific purpose for which you “optimized” that thing.
The above said and acknowledged, there are a number of features of the human body for which is it very difficult indeed to imagine any “optimization”-purpose that might have resulted in that feature ending up in the particular configuration it happens to be in. Examples: The esophagus, the human appendix, the recurrent laryngeal nerve, etc.
You are the poor schlepper who claims they don’t model evolution
Flint,
I agree with this. The question is to what degree are you convinced that the scientific hypothesis is real. As you can tell I am very skeptical that a trial and error process can create all life’s diversity. I think it can play a role but just that. There are two standards that are working here 1. Is the scientific method. This is where you need a repeatable process so you can make a prediction. I have used this process dozens of times and have a lot of confidence in it. 2. Inference to the best explanation: This is the standard used for evolution. I do not have a lot of confidence in this standard especially if there are very limited competing hypothesis. I think there is lots of data that show that evolution has occurred and all life shares common biochemical mechanisms. I think the answer to the cause of these changes is we don’t know yet. The fact that we cannot produce a GA that creates a novel protein with out a target is telling.
The problem with the competing hypothesis ID is that it only can use the inference standard because it is not offering a tangible mechanism. I feel the same with Michael Denton’s structuralism argument.
The fact is that we cannot design proteins. Nor is there any conceptual scheme in sight that wpuld enable designing biological molecules without trial an error. Directed evolution is the gold standatd in pharmaceutical research.
The large numbers that IDists are so fond of preclude having a database of functional sequences. The situation is worse for regulatory sequences.
There is no known way to regularize the principles of genomic sequences. No grammar or syntax. There are known modules that get mixed and matched, but no way to design a completely novel protein. Even something as simple as Lenski’s two or three step citrate metabolic pathway is beyond the ability of human design, except by trial and error.
It’s evolution or the highway. If IDists disagree, prove me wrong.
Seems this skepticism is quite common. It seems incredible that random dart-throwing plus selection could, given time, produce the degree of diversity we see. There MUST be something else going on, something we haven’t found yet, right? Yet whenever we take what we know is going on and run a simulation, we see enormous diversity appearing with amazing speed, from the simple expedient of (usually) retaining what works and (usually) discarding what does not.
The problem with the concept of “trial and error” is that it seems to forget BOTH the fact that each series of trials starts with what worked last series, and that this has been occurring for a period of time that simply boggles the mind. Nobody but nobody can conceive of billions of years.
Think, maybe, of the example of driving cross country. You don’t have a map, you don’t have any particular destination other than “west”, all you have is a compass. The pattern of roads (in the US) is fiendishly complex – they range from dirt tracks to superhighways, they go in all directions, some go in circles. With sheer trial and error, you have a “drunkard’s walk”, and you’ll never get there. But with a compass, you can retain the segments of your trip that moved you westward, and not take segments that have no westerly component. Your trip might take a long time, you might get sidetracked into some circles, you won’t set any land speed records. But you WILL get there, simply because selection works.
Without selection, trial and error will get you nowhere much.
I thought you were talking about GA’s.
Tell petrushka that.
I don’t have any issue with an evolution that is fine tuned to solve specific problems. like GA’s.
What do you suppose the word error implies.
It could be called trials and differential success, but I’ve been told that pedantry is frowned on.
Right. And I decided not to be a pedant about it and kept my mouth shut because it really didn’t much change the point I was making. But thank you for including the haploid case. Could you explain the disappearance of Ne in the haploid case?
Some suggestions for turning our little weasels into diploid digital organisms?
Thanks Joe.
Congratulations. You actually manged to get something right.
I did not know that your WEASEL program employed a near infinite s, but now that I do know that, it’s just another affirmation of my claim that GA’s do not model or simulate evolution and that yours certainly does not do so. [You agreed with that, didn’t you?]
Have you changed your mind now that Joe F has spoken? Is your GA now a model or simulation of evolution?
I think he was saying that you don’t understand the concept of selection. From your posting history, I assumed otherwise.
Or maybe he was trying to say that in emphasizing the power of selection, I was rejecting the idea of trial and error. Hard to tell, as usual.
The implication is that in Mung’s mind, you can have EITHER trial and error, OR selection, but not both. But it’s very hard to guess what he’s trying to say.
One of my weasel implementations uses crossover. Sexy mustalids, imagine that.
We all know that if you incant the right magic words, you win. It’s all about what words you use.