Look who one of the co-authors is in this journal abstract. 🙂

You can find the abstract HERE
Please show a little class in the comment section in consideration of my co-authors who are only guilty by association with me. You can trash talk about me, but in deference to the innocent parties, please don’t demean or trivialize their work, ok? Set a good example and celebrate what other people do in contributing to the advancement of knowledge.
Anyway, the abstract:
Topoisomerase II is a critical enzyme involved in unknotting and detangling DNA during replication, transcription, and cell division. Humans have two isoforms of topoisomerase II, α (Top2A) and β (Top2B), originating from genes on separate chromosomes and displaying distinct functional roles. In addition, these enzymes are the target of several successful anticancer therapeutics. Unfortunately, these agents are nonselective and a growing body of evidence implicates Top2B as a mediator of off-target toxicities, while Top2A is likely a better target for disruption of cancer cell growth. The isoforms share about 65.2% amino acid identity primarily in the N-terminus and the core regions, which contain the catalytic domains of the enzyme and the regions targeted by all clinically-relevant anticancer agents. On the other hand, the C-termini of the human enzymes share only ~30% amino acid identity across ~400 amino acids. The carboxy terminus does not participate in catalysis, but has been hypothesized to have a role in the regulation of topoisomerase II activity, which may explain how these proteins are independently regulated. Since the C-terminal region has been largely unexplored, we undertook an analysis to identify key differences between the C-termini that may help explain the differential regulation of the topoisomerase II isoforms. The relative rates of synonymous (dS) and nonsynonymous (dN) DNA substitutions between the isoforms suggests that these sequences are under purifying selection for functional constraint (dN/dS = 0.25; P < 0.0001, Z test, 1,000 bootstrap replicates). The 434 C-terminal codons, however, had a relatively high dN/dS ratio of 0.59 (P < 0.0001), reflecting elevated amino acid diversity. In addition, bioinformatic evidence from Phosphosite (Phosphosite.org) indicates that nearly half (91/191 for Top2A) of the putative post-translational modification (PTM) sites are found in the C-terminus. Of the PTM sites found in the Top2A C-terminus, over half (~50) are distinct from those found in Top2B. Aside from sequence characteristics, protein-protein interaction data from the Biogrid database (thebiogrid.org) indicate that ~143 proteins have interaction evidence with either Top2A or TOP2B. Of these proteins, only ~34 are confirmed to interact with both isoforms and several are known to interact with the C-terminal domain of Top2A or Top2B. Taken together, these data suggest distinct sequence, PTM, and interaction profile characteristics for the C-termini of the isoforms of Top2, which may provide critical insight into the differential regulation of these enzymes. We hypothesize that these results provide the foundation for topoisomerase II isoform-specific targeting strategies for anticancer therapeutics.
The highlighted portion above was my focus along with some of the co-authors. My involvement actually originated from a discussion I started at TSZ, specifically comments like this:
Promiscuous Domains are Better Explained by Common Design, Comment 8/29/18
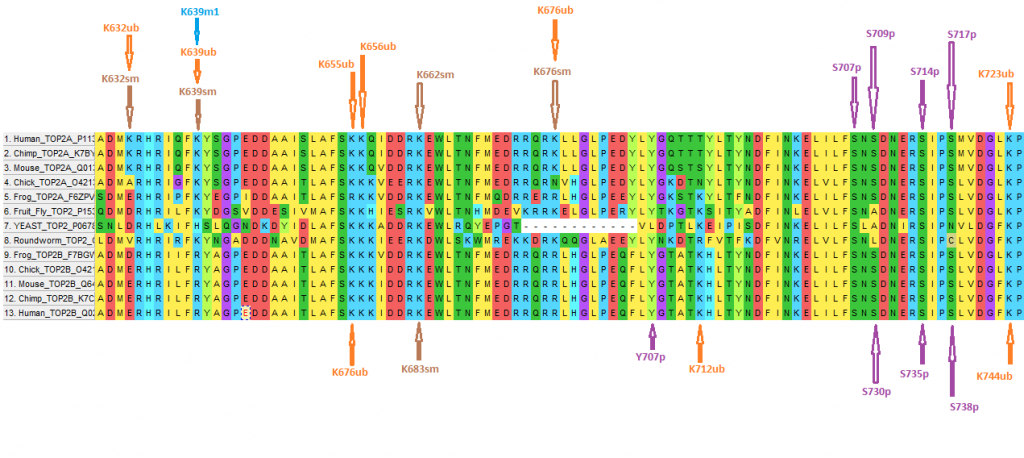
I mailed some of the graphs I developed and posted at TSZ to the lead author in September 2018, and he was so appreciative he listed me as co-author and used my graphs at a poster presentation.
It actually took about 4 weeks of combing through data to put something coherent since I was learning the ropes on the relevant databases, and there were several dead-ends in trying to connect the bio-grid data (mentioned above) to the PTMs I was cataloging. I was hoping to find promiscuous motifs in the proteins listed in the interactome/biogrid data with the motifs that were surrounding the PTMs in TOP2A and TOP2B, but to no avail since kinases perform the “write” operation on amino acids have degeneracy in what amino acid motifs they target.
This dead-end was so daunting, I had to simply quit and hope that work on Kirk Durston’s K-Modes algorithm could solve the problem of connecting the PTMs on TOP2A and TOP2B with the PTMs on the proteins listed in respective interactomes listed through biogrid. I mentioned Kirk’s paper HERE.
Again, if you wish to trash talk about me, I’ll accept it, but please don’t demean the hard work of my co-authors just because I’m listed with them (somewhat by accident).
Thanks again to all the participants at TSZ who help get my work showcased in the FASEB journal abstract and at the Conference of Experimental Biology this past April, 2019 connected to the publication of the abstract. The editorial review I received here at TSZ was crucial in helping prepare a compelling case for some of the ideas presented in the abstract and poster.

Try the entirety of Young Earth Creationism, or geocentrism, to pick just two major topics. Especially the former, but also to some degree the latter, is motivated by scriptural interpretations.
“By definition, no apparent, perceived, or claimed evidence in any field, including history and chronology, can be valid if it contradicts the scriptural record.” – Answers In Genesis
You will find no equivalent to that statement output by any secular scientific organization.
There’s also a branch of anthropogenic climate change denial motivated by Christian interpretations of scripture that imply God wouldn’t allow it because mankind were given dominion over the Earth and such nonsense.
Congrats Salvador! Great work getting published. Not a small feat, in my experience.
Thanks Eric, more stuff in the pipeline too. I’ll talk offline with you about it sometime soon. 🙂
Slight correction: an abstract doesn’t count as “getting published”, even if it’s in a scientific journal. Wait for the paper, if there ever is one.
And if it does get published, John Harshman is going to be the first one to congratulate Sal, not to mention apologizing to him…