One of the most brilliant evolutionary biologists of the present day, Richard Sternberg, PhD PhD was ousted and permanently blacklisted by the National Institutes of Health and the Smithsonian Museum for his ID sympathies.
Sternberg is neither a Creationist nor Darwinist but classifies himself as a Process Structuralist which means he is not much involved in the ultimate questions of how things came to be, he just appreciates the amazing patterns of similarity and diversity in biology.
He was labelled by some of his former supporters as an intellectual terrorist after he used his position as editor of a journal to publish an ID-friendly article by Stephen Meyer in 2004. He paid dearly for that decision, and his subsequent dismissal from the NIH and Smithsonian precipitated special investigations by members of Congress and the White House a decade ago. Unfortunately, nothing of consequence was done for Sternberg and he was destroyed professionally and personally.
Despite his circumstances, he continued to publish excellent essays such as the one that highlights the non-random patterns of SINES (presumed by some to be junkDNA) which are present in mice and rats (link below).
To understand his essay, I will describe the essentials of his essay with a parable. Suppose we had two mostly identical stories published. The stories are identical except for the fact that in one version of the story the name of the main character is “Mary” and in the other version, the name of the main character is “Caroline”. Even if the two versions of the stories were generated from the same template, the changes in the two descendant copies could not have been random.
So even if we assume some sort of common descent of the two versions of the story from some ancestral copy, the differences in the versions could not be the result of random copying errors, but very deliberate and methodical changes in the duplication process that created the two versions. This peculiar phenomenon plays out approximately in the genomes of mice and rats, and I call it the Sternberg-Collins paradox in honor of Sternberg who brought the paradox into prominence and Francis Collins who was among the first to comment on the anomaly.
Assume for the sake of argument rats and mice came from a common ancestor. Are there differences which are non-random and thus evidence for non-random mutation? Sternberg effectively answers, “yes”.
The mouse and rat genomes look very similar, but there are sequences that repeat over and over again in each of their respective genomes and mostly in the same corresponding locations. If these sequences were identical in both lineages, there would not be much of an issue, but they are different even though they are in the same general corresponding locations.
Let me call one set of these repeating sequences in the mouse genome “Mary” and the corresponding sequence in the rat genome “Caroline”. The name “Mary” appears in numerous places in the mouse genome, and in the corresponding places where “Mary” appears in the mouse genome, “Caroline” appears in the rat genome.
Of course this was a figurative way of describing what is going on. “Mary” is in reality the B1/B2/B4 set of SINE retro elements in mice, and “Caroline” is in reality the ID SINE retro elements in the mouse. But don’t get hung up on the fancy language, the basic problem of non-random changes from a supposed common ancestor is brutally evident. A graph that shows the non-random changes is here and explained in Sternberg’s essay:
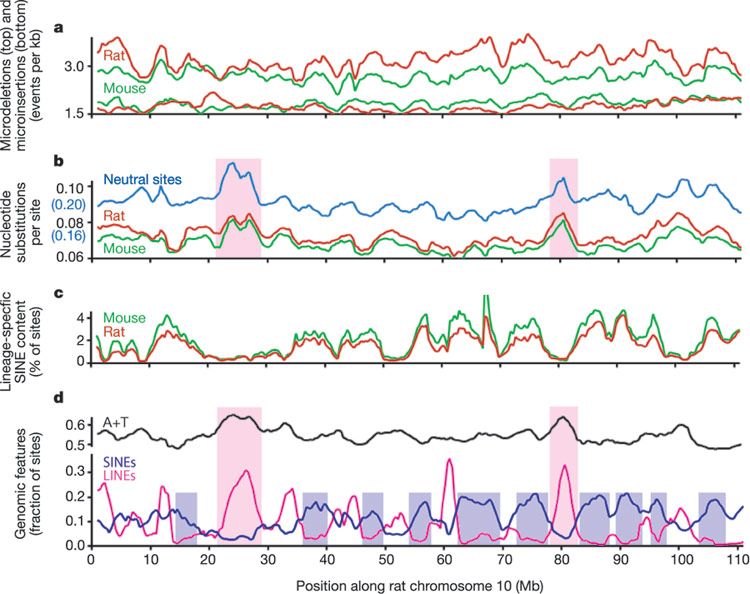
Sternberg’s point is that if mutational events happened, it was non-random, since to suppose it was random is an absurdity in the extreme. It cannot be the result of random DNA copying errors, but some non-random copying mechanism if common descent were true.
Is the non-random pattern the result of natural selection? Sternberg doesn’t address that question. But if the non-random SINE pattern is the result of selection, then this would mean the SINES aren’t junkDNA.
But if natural selection is assumed as the mechanism, there would be issues of the evolvabilty of so many nucleotides simultaneously. We’re talking maybe 300,000 SINE insertions in each lineage! For mice, the B1 sine is about 150bp and homologous to the 300bp primate Alu. The B2 SINE is 190bp. I could not find the size of the rat ID SINE nor the mouse B4 SINE, but I presume they are within the range of most other SINES (75-400 bp).
The source article in Nature which Sternberg referenced and had a buzzillion co-authors can be found here:
Genome sequence of the Brown Norway rat yields insights into mammalian evolution
That article points out:
Despite the different fates of SINE families, the number of SINEs inserted after speciation in each lineage is remarkably similar: approx300,000 copies.
Sternberg highlights the issue of having 300,000 non-random insertions happening in parallel in the two lineages after they split. He lays out the paradox which is the focus of this OP here:
Beginning to Decipher the SINE Signal
FWIW, I suspect there are probably similar issues with the Alu elements that appear in primates. Certainly this would be an issue if the mouse B1 (homologous to primate Alu) is in homologous locations in the primate genome. If that is the case, the Sternberg-Collins paradox is in play as well for primates. Perhaps one day we’ll know for sure if this is the case.

swamidass,
So do you mean the purpose of evolution was to evolve in a specific way such that inevitably organisms would evolve towards humans? That’s an interesting position.
I am not sure how that position differs very much from many so called creationists theory of life. Do you have a theory about the mechanism by which life strives towards evolving into humans?
I am slightly concerned that you have just lost the support of 98% of the other evolutionists here though. But that’s ok, they are a sensitive bunch.
phoodoo,
Yawn. Yes, I’m shitting myself over the lack of A Plan and the role of Chance.
I hope new people that come here read phoodoo’s posts. One barely needs to respond. I could think of very few things that would more effectively undermine whatever case I wished to defend than have phoodoo advocate for it.
I’m sorry if I wasn’t clear. I was just saying that it doesn’t follow from
We don’t fully understand how X happens.
that
God made X happen.
But that’s the leap you always make. We didn’t use to know how snowlflakes happened. Now we do.
ETA: first I wrote ‘didn’t used to know’; then I checked around, and while they’re both apparently ok ‘didn’t use to know’ is more common, so I changed it. But as they’re both awful, maybe ‘used not to know’ would have been best.
#Englishsucks
swamidass,
I would disagree. Darwin was describing selection dominated adaptive change, which is NS by definition. His theory certainly didn’t hinge upon such being the sole, nor even the main, mechanism of change in operation, such that he could be ‘falsified’ simply by its being relegated. He specifically mentioned the possibility of neutral change.
phoodoo,
Dr Swamidass appears, as best I can tell, to be able to keep religion and science separate in his scientific work. As such he has not ‘lost’ me as yet.There are plenty of theistic scientists producing good work out there.
That’s a problem with the marijuana relegalization movement as well. There are some proponents who set it back every time they manage to get interviewed. My only response in those cases is “An idea is not responsible for the people who hold it.”
I think I’m more of a theistic evolutionist than Dr. Swamidass!
OMagain,
Well, don’t rush to judgement just yet O’Magain, we still haven’t heard how this process that God started is supposed to inevitably end up with humans.
So you might want to wait. Your hatred of God can’t be disregarded that easily.
walto,
But Walto I still don’t understand. In the conversations you are referring to, it is about how to reconcile a belief in God, with an unguided process that could have made anything, it didn’t have to make man. So what in the world does, “we don’t know how it happened, doesn’t mean God did it” talk have anything whatsoever to do with this conversation.
God is already being assumed in this case, so what is your point?
what was that book? The ‘motes’ and ‘beams’ bit.
Mung,
I think after reading his description, its possible EVERYONE on this site is more of a theistic evolutionist than Swamidass.
But I guess I will wait for him to elaborate.
phoodoo:
Phoodoo apparently thinks OMagain is an Irish name.
ETA: And that “Wale’s” is the possessive of “Wales”:
Once again their obsession with leaders fails them. The entire of Wikipedia must be wrong because it’s leader does not worship at the feet of the same god as them, apparently.
What’s wrong with the Jesus page at wikipedia phoodoo?
https://en.wikipedia.org/wiki/Jesus
Why would an atheist preacher have a page on ‘their’ site about Jesus where he does not simply say it’s all nonsense? Phoodoo?
Dr. Swamidass,
I’m very grateful for you participation in this discussion. I think you are correct to raise the point about chemical affinities which may create preferential insertions of SINES even though I think your original suggestion of transposases (appropriate for Class 2 Transposable Elements) should be replaced with the mechanism of reverse transcriptases (appropratie for Class 1 Transposable Elements). Though I don’t think that is the mechanism, it can’t be formally ruled out as a possibility at this time, so as I said, I’m indebted to you for reinforcing my concerns. Though I remain skeptical that it evolved by that means, it would be appropriate to convey your concerns as I’m sure it will be also on the minds of others.
Theisitic evolutionists like yourself, John Polkinhorne, and Francis Collins were influential to sustaining the Christian faith I profess, but I arrived at my present viewpoints on creation for reasons similar to those who are present day ID proponents and creationists but were formerly atheists or theistic evolutionists — individuals like Dean Kenyon, John Sanford, Richard Lumsden. I myself was theistic evolutionist, then an old earth creationist, now I’m a Young Life Creationist.
Being a student of biology I do see the similarities of chimps and humans in the genome browsers, so I understand how powerful the similarities are. They are undeniable.
There may not be many areas we agree on regarding evolution, but I would simply put on the table maybe one area I hope we can agree on, namely, that the level functionality of the genome is not yet settled.
There are evolutionists like those in the ENCODE/RoadmapEpigenomics consortium (and perhaps by way of extension even the overseeing Director over the ENCODE/RoadmapEpigenomics consortium Francis Collins himself) that suspect the genome has a great deal of functionality.
Sternberg has used arguments such as the one in the OP to suggest SINES in mice and rats, and by way of extension Alu SINES in humans, are functional. The method of his deduction may or may not be valid, but it is formally possible his ultimate conclusion of functionality is correct.
Though, as Dr. Ayala rightly pointed out, Alu insertions in newborns in the present day are damaging, it is also true neuroscientists and genomic researchers seriously consider the Alu’s conserved in the genome are functional. This leads of course to a paradox. How can Alu’s (from the present day) be damaging but Alu’s (from the past) be so amazingly functional?
I think paradoxes such as these should not hinder consideration that the Alu’s have function in a way that is medically significant. The same could be said of many elements of the genome where there is contentious debate about its biological significance, especially in light of issues like the c-value paradox.
For myself, although the genetic code is nearly universal, I think the genomes are utilized in different ways in each organism from the regulatory mechanisms, to the use of chromatin modifications and the use of non-coding RNAs.
So to the extent that some at BioLogos promote the notion that major sections of the genome are junkDNA (like Dr. Ayala’s criticism of the Alu’s), I would be cautious because the sentiment at the NIH (which Dr. Collins heads) is that the genome is mostly functional.
The official figure of functionality is variable, but pretty much every lab worker I meet be they faculty or post docs in my classes or people in the conferences I attend have the strong sentiment the genome is highly functional.
Whether that turns out to be the case remains to be seen, but I think it’s a little premature to declare major sections of the genome as definitely junk from introns, to pseudogenes, to lncRNAs, to repetitive elements, etc. The private sentiment among the medical researchers at the NIH is that the genome has a lot of function. That sentiment does not arise from metaphysical considerations but the simple fact so many heritable diseases and cancers and other health issues they study seem to be highly correlated with genomic variation in the non-coding regions.
Finally, as an aside, I’ve occasionally attended functions at Dr. Collins church as he is my denomination.
I wish him and you and my brothers at BioLogos the very best. God bless you.
Sal,
As I said above, to the extent you are criticizing theists here, I’ll let them defend themselves. But your arguments from ignorance animate pretty much every thread you participate in. It’s kind of your M.O.
I have an enormous pool of ignorance to draw on, so I can make these arguments indefinitely. 😀
Once again, whether you notice it or not, you have equated “non-coding” with “presumed by some to be junk”. Please stop that. The genome has a lot of function even if only 10% of it isn’t junk, so that’s not a useful claim. Mutations in junk DNA can cause disease if they result in a new “function”, for example a new transcription factor binding site, in the wrong place.
One might ask why, if most of the genome is functional, very little of it is conserved over evolutionary time. I realize this is a problematic question because you don’t think evolutionary time exists. But is there an analog of the question that would make sense to you?
Here’s one. Supposing that rats and mice (i.e. Rattus and Mus) were separately created more or less as we see them, why should they have different retroelements in similar places? Why not the same ones?
No I have not, you’re misreading what I said. If people view the ncDNA regions as functional, then that includes the ncDNAs that were presumed as junk by haters of ENCODE, not just the ncDNAs like tRNAs. Do I have to draw a Venn diagram for you to help you understand the subtlety? 🙄
I was also specific about certain classes on ncDNA : introns, Alu SINES and the rest of the SINES, LINEs, introns, pseudognees, lncRNAs. Those are viewed by the haters of ENCODE as junk.
John, to Sal:
Because God likes variety.
And if they had been the same, my answer would have been:
Because God likes consistency.
I think I’m getting the hang of Creationist Logic™.
How about them ‘moebas, Sal?
Keiths,
I address the issue about ameobas indirectly. Did you bother studying the Robert Tjian videos I suggested Evolution Visualized thread regarding non-homology in regulatory networks. Genomes aren’t just about coding, but regulation and development (for multicellular organisms).
But first the amoeba:
http://salamander.uky.edu/srvoss/425SP08/Taftetal.pdf
and
You can see for yourself an amoeba does some things that are unique to its genome which humans don’t do with theirs: https://www.sciencedaily.com/releases/2005/02/050224115355.htm
And RNAs that are generated from repetitive DNAs can be altered and we’re only beginning to scratch the surface of post-transcription RNA modifications. Some call it the Epitranscriptome, and the NIH has 205 million dollar project in the planning stages just to study this stuff.
You’re citing arguments that go on the bad extrapolation of Monod’s famous claim: “What’s true for E. Coli is true for Elephants”. Well, E. Coli don’t process their genomes like Elephants do they?
Even identical genomes in the human cell types have different regulatory networks, and thus use the genome differently in each cell type. Tjian’s video illustrates this. You have to look carefully at the different deployment of distal activators for starters…
Thus if regulatory they lack homology between cell types in the same species, how much more between species!
Plants for example use their chromatin (of which DNA is a part) in ways humans don’t and vice versa.
http://pcp.oxfordjournals.org/content/55/11/1859.full
The chromatin memory can’t be expanded if there isn’t a corresponding change in genome size, even in the same species!
And what if distal activators and molecular machine binding sites are used differently for different classes of molecular machines.
You’re mistaken to think the genomes are used in the same way in each organism, and they aren’t even used exactly the same way between cell types.
A nice faceplant by Sal.
I posted a tweet that mentions an amoeba with a genome size of 670 Gbp, 200 times as big as the human genome.
Sal wants to argue that most of that amoeba’s genome is functional, so what does he do? He cites an article about E. histolytica, another amoeba.
The size of E. histolytica‘s genome? 24 Mbp.
That’s right. Sal is trying to use E. histolytica, an amoeba that gets by on a mere 24 Mbp, to support his ideological claim that most of P. dubium‘s 670 Gbp are functional.
Oops.
To put that in perspective, P. dubium‘s genome is more than 27,900 times as large as E. histolytica‘s.
Perhaps next time Sal will do more reading and less Googling, copying, and pasting.
Now, that’s substantive! 🙂
What’s an error of just 0.068 dembskis between friends?
walto,
Pointing out that in the discussion we are having, God is already assumed is an argument from ignorance? Let me think about that? Hm,…well, that’s insane Walto!
There is no argument of ignorance here, other than the argument that you are calling an argument of ignorance, which is an ignorant argument.
Your are back to your pointless speeches which aren’t even remotely related to the discussion at hand. Actually, its not even a speech as much as it is your constant wailing reminder, “I HATE GOD.”
Yea, well, sorry.
I was debating someone online who advocate the idea most of the genome is junk. He kept arguing the C-value paradox.
I said in response that if even cell-types in same species use the genomes in different ways, then it stands to reason different species use their genomes differently and can thus have different sizes. I pointed out the fact each human cell type probably has a different regulatory network than another cell type.
He disputed that point.
I responded with this diagram of Genome-wide survey of tissue-specific microRNA and transcription factor regulatory networks in 12 tissues from:
http://www.nature.com/articles/srep05150/figures/6
Bwahaha!
Sal quickly changes the subject.
Gee, I wonder why?
Wait, you mean that different genes are active in different cell types in the same body? Whoa, who knew? And of course from this follows Jesus.
I think I’m getting the hang of this:
1. P. dubium has 2,970,000 percent more base pairs than E. histolytica.
2. Apply Creationist Logic™.
3. Therefore, P. dubium‘s genome is mostly functional.
And:
1. Gene expression and regulatory patterns differ by cell type in a body.
2. Apply Creationist Logic™.
3. Therefore, the genome is mostly functional.
And:
1. The genome is mostly functional, based on Creationist Logic™.
2. Apply still more Creationist Logic™.
3. Therefore, Jesus.
How am I doing, Sal?
This is powerful evidence of how differently they utilize their genomes to do business. 🙂 I provided papers that show the research of the unique usage of genomes of amoeba (P. dubium is an amoeba).
Differences of how genomes do business is powerful evidence against evolution.
The protein coding genes may be similar, but not the non-coding regions that contain recipes to create stuff like that depicted below — for example plants don’t have non-coding DNAs like Alus.
10%-11% of the human genome has 1 – 1.5 million primate specific recipes called Alu ‘s.
These Alus were considered excess junk of no utility, but now we realize it was pretty important and is what sets primates apart. In similar manner, those large amounts of DNA in other creatures can be used to do business differently in other organisms. Maybe in time we’ll see, but one thing we know, evolutionary theory didn’t help us figure stuff like the diagrams depicted below, molecular biology and the ENCODE consortium did.
Regarding p dubium, the genome size of half a trillion should be treated with caution as mentioned earlier (which Keiths ignored).
That said:
http://www.science.smith.edu/departments/Biology/lkatz/documents/McGrath_Katz_Tree.pdf
The genomes have widely varying sizes between and within species liekly because they do business differently with their genomes. Why do evolutionary biologists have such a hard time understanding this simple concept? I guess they don’t like the fact such diversity is evidence against evolution and certainly makes a mess of their attempts at phylogenetic story telling.
The diagram shows connections to a class of ncDNAs that create the microRNAs in the diagram.
This shows the principle that much of the recipe building involves non-coding DNAs in genomes, and these are involved in creating the regulatory networks. It is these non-coding DNA/RNAs that are often implicated in the vast differences in genome size between and within species.
Rgarding the ameoba genome, in one article I cited, it said:
tRNA genes code tRNAs which are non-coding, and further tRNA genes constitute 10% of the amoeba genome. Btw, tRNA’s can have large numbers of post-transcriptional modifications. 🙂
http://rnajournal.cshlp.org/content/21/4/642.full
Post transcriptional modification of tRNAs is a means of regulation. The amoeba appears to have a different regulatory mechanism and thus has a different genome to implement it that uses lots of ncDNAs to do the job!
https://en.wikipedia.org/wiki/Transfer_RNA#tRNA_genes
Sal,
For someone who likes to think of himself as a skilled gambler, you sure don’t know when to fold.
See what you’re doing there? You take one class of RNAs, a few of which are shown to be functional, and extend that to all non-protein-coding DNA, everywhere. So why aren’t most of them conserved in sequence between closely related species?
And you’re still talking about the wrong amoeba too.
I have one more question. If 10% of the human genome isn’t enough to build a human, or so you intuit, does your intuition tell you it should be enough to build a fugu?