Let me clarify that by saying that I mean a theory that purports to be a scientific theory, that makes testable predictions (“if ID was correct we should observe…, if ID were not correct, we should not observe…”) and is an alternative explanation for the diversity and extent of the pattern of life we see on Earth.
This is far from the first time I’ve asked this question. A quick search finds a couple of threads at Uncommon descent but I’ve had no satisfactory replies beyond the usual default that says something like:
The theory of intelligent design holds that certain features of the universe and of living things are best explained by an intelligent cause, not an undirected process such as natural selection. Through the study and analysis of a system’s components, a design theorist is able to determine whether various natural structures are the product of chance, natural law, intelligent design, or some combination thereof. Such research is conducted by observing the types of information produced when intelligent agents act. Scientists then seek to find objects which have those same types of informational properties which we commonly know come from intelligence. Intelligent design has applied these scientific methods to detect design in irreducibly complex biological structures, the complex and specified information content in DNA, the life-sustaining physical architecture of the universe, and the geologically rapid origin of biological diversity in the fossil record during the Cambrian explosion approximately 530 million years ago.
I wonder if any of our contributors can finally enlighten me as to what the scientific, biological theory of “Intelligent Design” actually entails.

CSI is not a measure that tells you how “full of design” something is. I am working on a measure though to tell how full of shit someone is. Interested?
STRAWMAN!!!
Mine’s bigger than yours.
Wrong and wrong again. (I just felt like saying that.)
I apologize for my contributions to that confusion.
Exactly the point I was trying to get across to Frankie before I gave up.
Have you considered investing in a brick wall?
What Vincent said is that we shouldn’t expect more than three top level systems in the hierarchy. That seems completely arbitrary to me.
I don’t see why that would be a prediction of evolutionary theory. What we would expect to see is that those systems appear in a tree-like fashion
No, he said we should’t expect to see the same top level limitation of 3 systems, that is, any number of top level systems work for ID. because ID is compatible with any and every arrange of those systems and their hierarchy: the theory doesn’t predict anything. It just happened, that’s all, it isn’t a theory.
That’s not true. I said that, for the sake of argument, let’s say that Vincent’s criteria to falsify evolution is valid, and the criteria is met by the evidence.
That would mean that evolution can’t explain the observed complexity. The rest of the argument works the same
Well, that’s wrong. He says: “I’d expect the number to decline exponentially for higher levels” and he throws some numbers out there for bacteria and eukaryotes, all completely arbitrary IMO, but again, it doesn’t matter at all
Both CSI (biology) and FSC refer to the complex sequence specificity required to produce a function. Read Dembski and Meyer- both make it clear that CSI (biology) relates to sequence specificity that produces a function. Biological CSI has always pertained to function derived from the sequence specificity.
Perhaps you guys have never read “No Free Lunch”, “The Design Revolution” and “Signature in the Cell”. All three books make it clear that CSI (biology) is FSC.
No Joe, I am saying that Durston’s concept of FSC is the same as Dembski/ Meyer’s concept of CSI in biology.
One more thing: Extant bacteria, unicellular eukaryotes and multicellular eukaryotes are all equally “evolved”. Why should we expect to see different levels of depth in their complexity if evolution is true?
And he responded to you. He did a great job of explaining how sadly you were mistaken. I am not sure why you would want to dig that up again
I do agree that those specific numbers may be somewhat arbitrary but it seems to me the crux of his argument is that under Darwinian evolution we would expect a distribution, whereas if we do discover such a distribution “if life were designed, there would be no reason to expect that.”
So he has not put some arbitrary limit on evolution, beyond which he claims it cannot go. He doesn’t say that evolution can do 6 hierarchies but if there are ten it means design. And he has not said that if we don’t find such a distribution that this favors intelligent design.
You can certainly disagree that there would be any distribution at all under Darwinism, but it seems to me that we do in fact have something that could be tested. I found it an interesting proposal.
I wouldn’t call it an intelligent design prediction though, because I don’t see how the absence of such a distribution would support ID.
I would deny the premise that all taxonomic groups are “equally evolved.” For example, I might argue that based on population sizes, generation times, and mutation rates, that bacteria are the most highly evolved organisms on earth.
AFIAK there is no scientific metric for how “evolved” some organism is.
Dembski/ Meyer CSI (biology) is supposed to be a mathematical measure of functional information of the functional sequence complexity observed in protein family biosequences as well as their RNA and DNA antecedents
Durston et al posit their own mathematical measure of functional information, in units of Fits, of the functional sequence complexity observed in protein family biosequences has been designed and evaluated.
They are both measuring the same thing. If you have CSI (biology) it is because you have functional sequence complexity. And if you are measuring the FSC then you are measuring the CSI.
They’re all “equally evolved” in the sense that they’ve all been around for the same amount of time.
Some branches evolved more “functional coherence” than others
How is that not not putting some arbitrary limit on evolution, if you agree his figures are arbitrary?
His figures don’t specify any limit, therefore, while they may be arbitrary, they do not place some arbitrary limit on the least frequent level of hierarchy.
Maybe there is some organism that has a level 20 hierarchy, but it has only one level 20 hierarchy. How many systems with a lower level hierarchy does it have? What does the distribution look like?
Here’s a link to his original post:
Nowhere in that post does the word “limit” appear.
Mung,
If one affirms that if evolution is true, then we should expect X, you’re limiting evolution in the sense that if you don’t observe X, then evolution is falsified. It can’t be that hard.
If such “X” is based on arbitrary figures, not derived from any evolutionary tenets, then it’s meaningless BS
…and what applies to figures, also applies to “distributions”. It’s all made up nonsense. What’s the operational definition for levels of complexity here anyway?
Mung quoting VJT quoting Axe: “But if life were designed, there would be no reason to expect that. “
Without knowing a single thing about the Designer, its abilities, limitations, likes and dislikes, purpose for the design, etc. how did Axe determine that a certain configuration is what to expect or not expect if life was designed?
One last time.
VJT didn’t say that if we do not observe the expected distribution [under his hypothesis] that evolution is falsified.
does The Designer only do things we’d expect, or does it operate in, let’s say, ways that might seem, perhaps, mysterious?
Because if it’s the latter, then Axe has no point.
Will it be in ” Frankies “?
LOL! We measure error magnitude in “Dembskis”, we measure bullshit in “Frankies”.
Who says IDiots can’t make a valuable contribution to science? 🙂
Well, you know, that’s how science works. It’s not a matter of opinion, it’s simple logic:
https://en.wikipedia.org/wiki/Modus_tollens
Micro-frankies for day-to-day usage with most people.
Nope. Dembski, Meyer and Durston might all be using the same scale, sure. But the distributions used by Dembski (at least from 2005/2006 on, he claims he always used it that way) involve comparing what we observe to a distribution expected from the natural evolutionary mechanisms. He does not say how to come up with that,
Durston, by contrast, uses a random distribution with equal frequencies of all 20 amino acids at each site of the protein. That’s similar to what Hazen et al. do.
The two are not the same.
You are not even listening to my argument. You just ignored what I said and posted regardless. My argument has NOTHING to do with how FSC or CSI are measured.
Try again, this time try responding to my actual argument and not what you want me to say:
Dembski/ Meyer CSI (biology) is supposed to be a mathematical measure of functional information of the functional sequence complexity observed in protein family biosequences as well as their RNA and DNA antecedents
Durston et al posit their own mathematical measure of functional information, in units of Fits, of the functional sequence complexity observed in protein family biosequences has been designed and evaluated.
They are both measuring the same thing. If you have CSI (biology) it is because you have functional sequence complexity. And if you are measuring the FSC then you are measuring the CSI.
Frankie,
Well, I already pointed out Durston’s error. So that must mean, if you are correct and Dembski CSI (and Meyer, if you say so) are the same as Durston fits, they are wrong, too.
And Durston corrected you.
How did he do that? Can you link to where Durston addresses my point about functionality in unknown proteins?
ETA: I’ve looked through the comments again and see no refutation from Durston regarding the rarity of novel functional proteins in sequence space.
Joe Felsenstein,
Masterful.
Joe’s had a cry about this and you on his blog: http://intelligentreasoning.blogspot.com/2017/01/joe-felsenstein-and-patrick-may-very.html
Richardthughes,
Hey, Blipey’s back. Oops off-topic 🙁
So, your argument “has NOTHING to do with how FSC or CSI are measured”.
But “they are both measuring the same thing”.
So it makes no difference that the two measures use different distributions of “chance” sequences? And thus would get different numbers? Numbers that could not be converted into each other. They are nevertheless “measuring the same thing”?
Frankie Logic (TM)
I made my argument perfectly clear, Joe. And nothing of what I said pertains to how they are measured. Just because two different teams came up with two similar yet different methodologies to measure the same thing doesn’t mean they aren’t measuring the same thing nor does it mean that CSI (biology) is the same thing as FSC.
Address my argument. If you actually read what I say I make the case that CSI (biology) is the same as FSC.
Can you say how that is relevant? Can you show how blind and mindless process can produce any proteins without the aid of proteins?
The answer is no to both questions, Alan
Science doesn’t care about the unknown, Alan. Do you want to know why? Because it’s unknown and as such may not even exist.
Yes Alan, science only cares about what was never unknown.
Wow. So science never does any research to learn about previously unknown things. I’ve heard some gobsmackingly stupid Creationist claims before but that’s on a whole different level.
Of course it does explain why ID-Creation “scientists” never do any research or produce any positive results.
It’s pretty clear at this point the subject matter is beyond Joe.
To repeat, Durston’s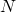 in the paper he linked to is defined as “the total number of possible sequences, both functional and non-functional.” As evolutionary processes do not (nor do they need to) perform an exhaustive search, using his
in the paper he linked to is defined as “the total number of possible sequences, both functional and non-functional.” As evolutionary processes do not (nor do they need to) perform an exhaustive search, using his 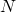 is not modelling evolutionary processes. He is beating up a strawman.
is not modelling evolutionary processes. He is beating up a strawman.
That sentence could be parsed in several ways or be taken as nonsense. You need to clarify rather than muddify.
Don’t ask questions if you don’t want answers!
This is not addressing the post.
Lion Fckface doesn’t care about the rules.
Alan Fox,
Didn’t read the post directly before yours, huh Alan?
You’re not listening. Not beyond, unknown.
This is false. I’ll be working on trying to come up with a simple enough way to explain it to you so that you’ll be able to understand it. Please be patient. Thank you.
You cannot show that blind and mindless processes are up to the task of producing proteins. And you can’t show that your alleged unknown proteins is anything but your imagination.
Can you show how blind and mindless process can produce any proteins without the aid of proteins?
Your response can be parsed in several ways or be taken as cowardice. Look Alan, you don’t have any evidence that blind and mindless processes can produce proteins from scratch.
Guess what functional sequence complexity pertains to? If you said “functionally specified sequential arrangements“, give yourself a cookie.
ID theory entails following :
Coded Information which is complex and instructional/specified found in epigenetic systems and genes, and irreducible , interdependent molecular machines and biosynthetic and metabolic pathways in biological systems point to a intelligent agent as best explanation of their setup and origins.
http://reasonandscience.heavenforum.org/t1659-confirmation-of-intelligent-design-predictions
Observation: Intelligent agents act frequently with an end goal in mind, constructing functional irreducibly complex multipart-machines, and make exquisitely integrated circuits that require a blueprint to build the object. Furthermore, Computers integrate software/hardware and store high levels of instructional complex coded information. In our experience, systems that either a)require or b)store large amounts of specified/instructed complex information such as codes and languages, and which are constructed in a interdependence of hard and software invariably originate from an intelligent source. No exception.
Hypothesis (Prediction): Natural structures will be found that contain many parts arranged in intricate patterns, metabolic pathways similar to electronic circuits, and irreducible structures that perform specific functions — indicating high levels of Information, irreducible complexity, and interdependence, like hard/software.
Experiment: Experimental investigations of DNA, epigenetic codes, and metabolic circuits indicate that biological molecular machines and factories ( Cells ) are full of information-rich, language-based codes and code/blueprint-based structures. Biologists have performed mutational sensitivity tests in proteins and determined that their amino acid sequences, in order to provide function, require highly instructional complex coded information stored in the Genome. Additionally, it has been found out, that cells require and use various epigenetic codes, namely Splicing Codes, Metabolic Codes, Signal Transduction Codes, Signal Integration Codes Histone Codes, Tubulin Codes, Sugar Codes , and The Glycomic Code. Furthermore, all kind of irreducible complex molecular machines and biosynthesis performing and metabolic pathways have been found, which could not keep their basic functions without a minimal number of parts and complex inter wined and interdependent structures. That indicates these biological machines and pathways had to emerge fully operational, all at once. A step wise evolutionary manner is not possible. Furthermore, knock out experiments of all components of the flagellum have shown that the flagellum is irreducible complex.
Conclusion: Unless someone can falsify the prediction, and point out a non-intelligent source of Information as found in the cell, the high levels of instructional complex coded information, irreducible complex and interdependent molecular systems and complex metabolic circuits and biosynthesis pathways, their origin is best explained by the action of an intelligent agent.